Novel Immunotherapy Targets Revealed by Genome-Wide CRISPR Screen
In a recent study published in the journal Cell, by Eric Shifrut, PhD, and colleagues from UC San Francisco and UC Berkeley, a powerful method for genome-wide knockout screening in primary T-cells was developed using the CRISPR-Cas9 system and combined with single-cell sequencing to reveal novel regulatory pathways and immunotherapy targets. In the development of this new method, oligonucleotide pools from Twist Bioscience were used as single guide RNAs and delivered into T-cells with the addition of Cas9 for gene specific editing.
It is well known that our immune systems detect and destroy foreign invaders like viruses and harmful bacteria. What’s less known is that the immune system has a heavy hand in protecting us against our own cells turning against us in the form of cancer. Immune cells constantly survey the body for precancerous cells and kill them when discovered. However, some precancerous cells evolve mechanisms to hide from the immune system, which allows them to continue growing uncontrollably and develop into a cancer.
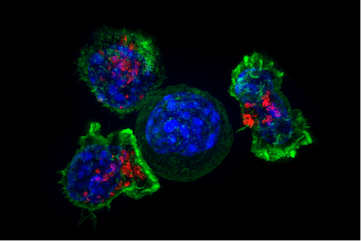
Superresolution image of a group of killer T-cells (green and red) surrounding a cancer cell (blue, center). When a killer T-cell makes contact with a target cell, the killer cell attaches and spreads over the dangerous target. The killer cell then uses special chemicals housed in vesicles (red) to deliver the killing blow. This event has thus been nicknamed “the kiss of death.” After the target cell is killed, the killer T-cells move on to find the next victim. (Source: From Wikimedia Commons)
There are several ways in which a cancer can avoid detection by the immune system, such as emission of chemicals that actively suppresses immune cell activation, or repression of signaling proteins required for immune detection. Recently, exciting successes have been achieved in cancer therapy by treating patients with therapies that bypass cancer blockades to activate the immune system. This new way of fighting cancer, called immunotherapy, has taken the field by storm, with several prominent pharmaceutical companies developing or already offering different forms of immunotherapy.
Although there are several immunotherapies currently on the market, they only help a small subset of cancer patients. Since no two cancers are the same, there need to be many therapeutic options available to free the immune system from the many cancer roadblocks.
In a recent study, published in the journal Cell, by Eric Shifrut, PhD, and colleagues from UC San Francisco and UC Berkeley, entitled “Genome-wide CRISPR Screens in Primary Human T Cells Reveal Key Regulators of Immune Function,” researchers uncovered the precise mechanisms necessary for immune activation, and revealed cellular molecules that have potential as novel therapeutic targets for the development of new immunotherapies.
To identify the critical players in immune cell activation, the researchers developed a cutting-edge screening method using CRISPR to edit genes on a genome-wide level in immune cells critical for cancer clearance, called T-cells. CRISPR editing takes place inside a cell when the Cas9 protein, which cuts DNA, is combined with a single guide RNA (sgRNA), which directs Cas9 to a specific DNA sequence. The researchers used a pool of 77,441 sgRNAs, each targeting a different DNA sequence, to effectively knock out a total of 19,114 genes. Lentiviruses carrying each sgRNA were created to introduce the sgRNAs into the T-cells, creating a population of T-cells with varied CRISPR targets in each cell.
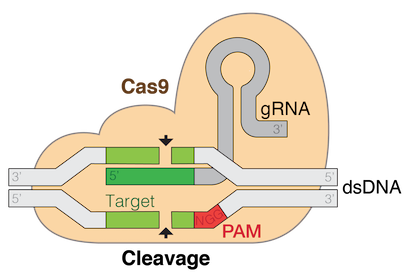
The Cas9 protein (tan shape) combined with guide RNA (gRNA, aka sgRNA) makes up a complex for targeted editing of genomic DNA, commonly referred to as CRISPR editing. (Source: Wikimedia Commons)
One major challenge of the screening experiment was working with T-cells elucidated directly from blood donors, which are notoriously difficult to work with due to their short lifespan in cell culture and low efficiency of Cas9 expression when delivered by lentivirus. To work around the low Cas9 expression, the researchers used a technique called electroporation, which uses an electric shock to deliver the Cas9 protein directly into the T-cells.
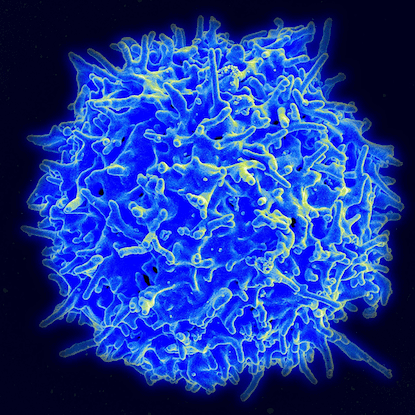
Scanning electron micrograph of a healthy human T-cell. (Source: Wikimedia Commons)
With Cas9 and sgRNAs combined within the T-cells, the CRISPR complex was able to make the desired gene edits within the population of cells. Each T-cell could contain several gene edits. The researchers then stimulated the edited T-cells with a chemical T-cell activator and, after a few days, they sorted the cells into non-proliferating and highly proliferating groups. The DNA in each group was separately isolated and sgRNA amplified, barcoded, and sequenced to reveal which genes inhibited or activated T-cell proliferation.
The screen successfully identified dozens of gene edits that positively and negatively modulate T-cell proliferation. The next step was to further verify that the genes identified in the screen did, in fact, have a role in T-cell regulation, using more stringent and targeted experiments. In these assays, the researchers focused on a set of negative regulators of proliferation that also exhibited a high therapeutic potential to enhance T-cell function when targeted. They found, for example, that when the genes UBASH3A, CBLB, CD5, and RASA2 (proteins involved in protein degradation, cell surface signaling, and GTPase activation) were knocked out, proliferation of T-cells stimulated with a chemical T-cell activator increased more than two-fold. Interestingly, these genes did not increase proliferation of untreated T-cells, indicating that they are not general regulations of proliferation, but instead modulate proliferation specifically induced by T-cell receptor signaling.
Taking their investigation a giant leap further, the researchers used CRISPR to knock out a small subset of the newly identified regulatory genes in T-cells, and used single-cell RNA sequencing to precisely characterize the cellular programs controlled by each gene. This analysis revealed unique cell states resulting as a consequence of the gene knockouts. The data also demonstrated that targeting certain negative regulators of proliferation can induce specialized gene programs that could potentially enhance the potency of T-cells.
Finally, the researchers tested their newly discovered T-cell regulatory genes in an in vitro cancer-killing assay. As predicted, knockout of LCP2 – identified in the screen as essential for a strong T-cell receptor stimulation response – robustly disabled T-cell killing of the cancer cells. In contrast, knockout of negative regulators TCEB2, SOCS1, CBLB, and RASA2 each significantly increased tumor cell clearance compared to the controls. These experiments served as important evidence that knocking out the function of these genes does, in fact, change the cancer-killing abilities of T-cells and presents highly probable immunotherapy targets.
This study has exciting implications for the development of novel, targeted immunotherapies. By revealing not only the genes involved in regulating T-cell activation and proliferation, but also the precise molecular pathways that are turned on when those genes are missing, the researchers have illustrated a complete picture of T-cell activation. The information unearthed can immediately be used for a more informed and comprehensive development of novel immunotherapies in the fight against cancer.
Featured image: T-cells attacking cancer cell (Source: Adobe Stock)
What did you think?
Like
Dislike
Love
Surprised
Interesting