Twist Bioscience Clonal Genes Aid New Cell-Free Protein Synthesis Technologies
In the field of protein biology, obtaining purified, soluble, correctly folded proteins is essential to research efforts. Typically, researchers use lab strains of bacteria and chromatography for the task. Bacteria are provided with new genetic material, encoding a protein of interest, and the bacteria’s own protein expression toolbox makes copies of the protein as it grows. These proteins are then harvested, purified by chromatography, concentrated, and stored ready for further experimentation. If a workflow for a single protein is highly optimized, the entire synthesis process takes between three and four days.
However, many proteins are difficult to express as they may fold poorly inside the lab strain of bacteria, aggregate together, or potentially cause harm to the bacteria, stopping its growth. These challenges mean that the life of a protein researcher is often spent optimizing and conducting protein expression instead of experimenting on the proteins themselves. If a company wants to make a chemical product using an enzyme, the difficulty in optimizing the synthesis of the enzyme to high volumes causes a trade-off against the potential commercial benefit the enzyme may bring. Ultimately, the current state of protein purification leads to unpredictable R&D pipelines.
Therefore, alternative strategies for protein synthesis outside of bacterial expression are desirable for the simpler production of useful proteins. Advances required in the field include more robust synthesis of difficult-to-express proteins, faster and/or cheaper roadmaps to obtaining the final pure proteins, or more rapid optimization cycles.
Cell-free protein expression is a promising tool toward this goal. Protein synthesis requires a number of key cellular components. These include the proteins that transcribe DNA to messenger RNA, ribosomes and tRNAs that translate messenger RNA into the sequence of amino acids in the protein, and cofactor molecules that all of the aforementioned components require to function. Live bacterial cells are convenient for protein expression as they can be grown cheaply and they not only contain but also constantly replenish their expression machinery. Cell free protein expression aims to replace the need for constant replenishment of machinery and thus the reliance on cell growth by saturating a solution with every part needed to make a given protein.
In a recent paper, published in ACS Synthetic Biology, Benjamin Des Soye and colleagues in the Jewett Lab at Northwestern University, reported their development of a brand new and simple cell-free expression system producing high protein yields. Their strategy was similar to other developed cell-free expression systems that used the extracted lysate (the complete set of proteins and molecules inside a cell) of E. coli bacteria, however, the researchers focused instead on lysate from the bacteria Vibrio natriegens. A microorganism that in nature makes its home in the salt marshes of Georgia, V. natrigens has recently become a promising tool for synthetic and molecular biology as it both grows twice as fast as E. coli, and can be easily supplied with new genetic material.
In the present study, the authors hypothesized that V. natrigens’ rapid growth rate required the microorganism to coordinate its cellular functions rapidly and thus the bacteria should have an inbuilt, highly efficient system for protein synthesis. It follows that lysate from this organism could provide a source of highly active protein translation machinery, potentially making it an ideal base for cell free protein synthesis. In concurrence with this hypothesis, the authors note that each V. natrigens cell contains almost 40% more ribosomes than a typical E. coli cell.
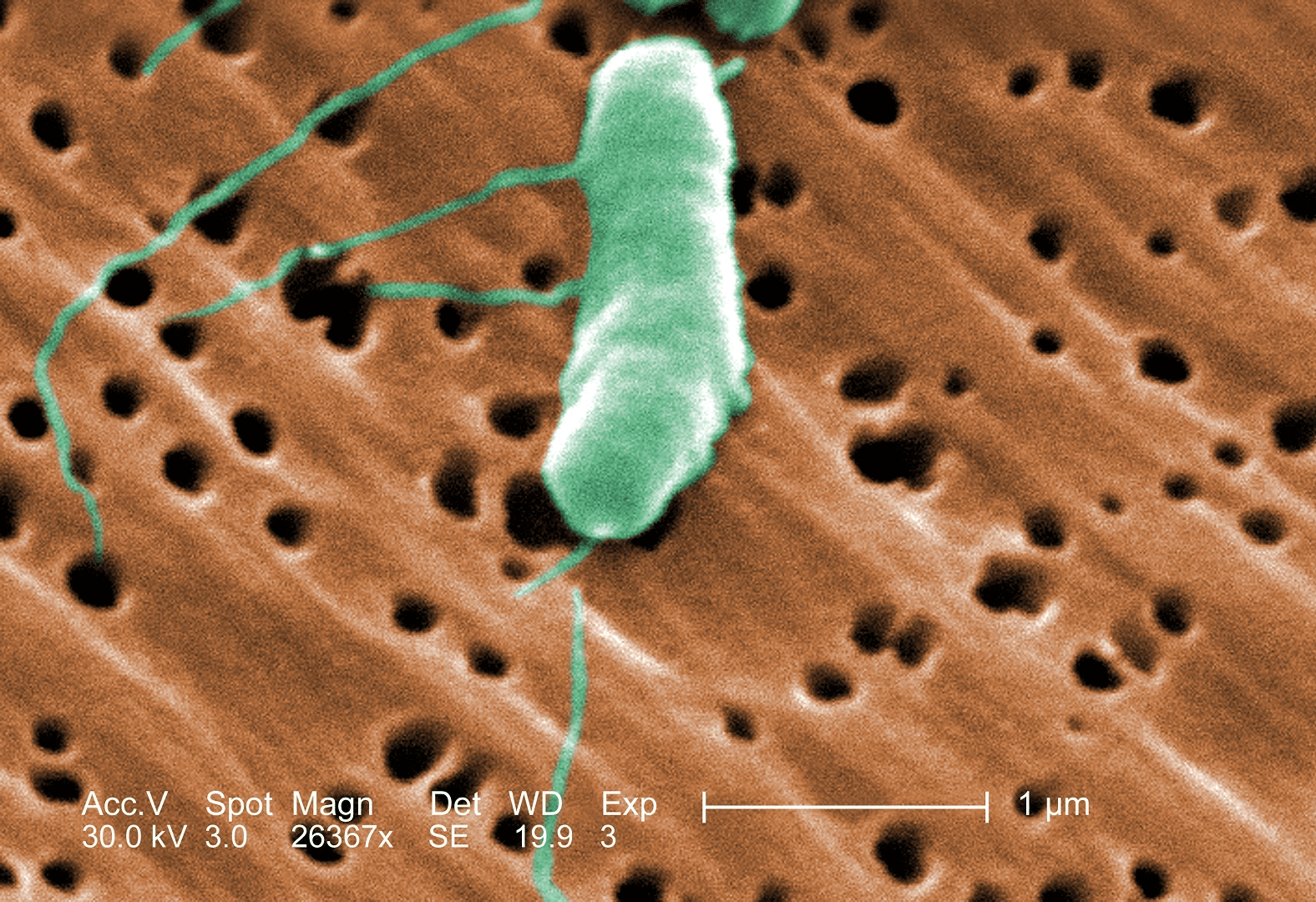
V. natrigens is from the bacterial genus Vibrio. These bacteria are all characterized by their curved-rod shape, and their flagella - the long green extensions in the above Electron microscope image. Flagella provide the bacteria the ability to swim. Public domain image source: Wikicommons.
The paper’s authors were able to develop favourable cell-free expression conditions from V. natrigens lysate by removing known cell-free expression inhibiting genes from the bacteria’s genome and optimizing its growth conditions. They then sonicated (blasted with high frequency sound energy) cultures of the bacteria to break open each cell, forming a lysate that was then supplemented with extra amino acids and energy providing molecules. With the resulting rather crude mixture the researchers achieved robust protein expression of fluorescent proteins when adding in plasmids containing their genes. From 1L of culture, high concentrations of 1.6 mg/ml protein were produced in as little as 20 hours. These yields are comparable to state-of-the-art E. coli-based systems that have been developed over the preceding 20 years, the key difference being that this proof-of-principle V. natrigens system still has plenty of room for optimization.
Importantly, the team used Twist Bioscience clonal genes for their expression experiments. One of the key benefits of cell-free protein synthesis is rapid turnaround times. Therefore, the creation of plasmids for protein expression presents as a potential bottleneck in system optimization. Traditionally, labs order synthetic gene parts and clone these into plasmids with standard molecular biology techniques. Using these methods, the timeline from DNA order to complete plasmid construction is between two weeks and even a couple of months in length. Instead, Twist Bioscience onboards plasmids designed by researchers. Then, Twist Bioscience delivers a desired gene, sequence verified, and cloned into a vector of the researcher’s choice in approximately 10-15 business days.
Additionally, Twist Bioscience’s high-throughput DNA synthesis technology provides cloned genes at market-leading prices, which are often less expensive than ordering the gene as a fragment from elsewhere. Combining Twist Bioscience clonal genes with the V. natrigens cell-free expression system provides a resource of rapid, low-cost protein expression, which could enable considerably increased throughput in protein research.
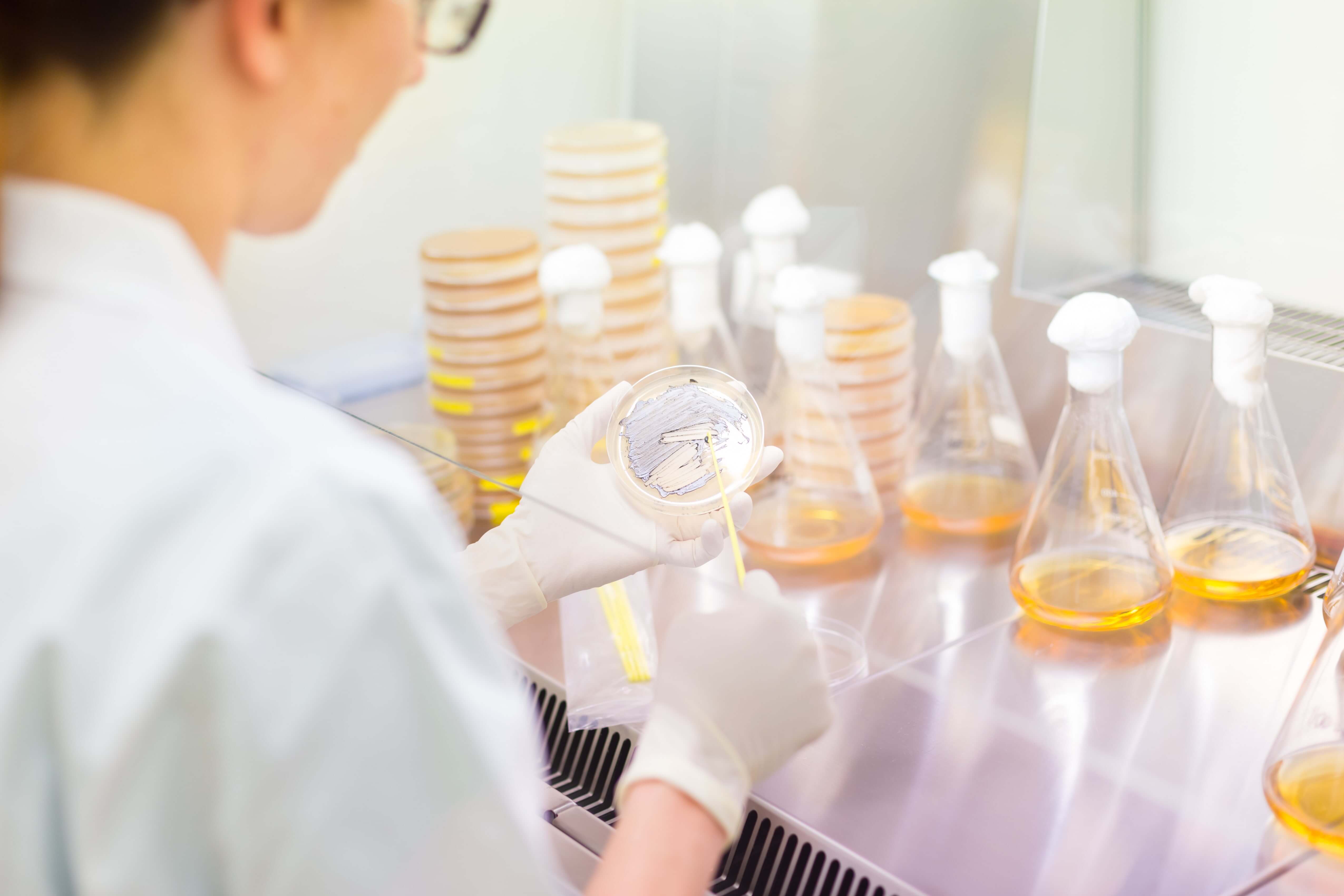

Traditional protein purification requires the lengthy culture of bacteria for every protein purified. Cell-free purification methods disconnect growth and purification, as pre-prepared mixtures from cell lysate are used to express and purify proteins without the need for cell growth. Source: https://stock.adobe.com/images/life-science-researcher-grafting-bacteria/84996645
In their further testing of the system, the researchers also showed it possible to lyophilize (completely dry) their lysates in a dehydroprotectant compound, and rehydrate them after a week’s storage without losing expression efficiency. Expression lysate can therefore be batch produced and stored easily for future use, significantly shortening the time-to-protein of any subsequent protein expression experiments. If a researcher has their desired protein encoding plasmid in stock already, their time-to-protein can potentially become as little as a day.
Finally, to show off their cell free system’s potential application, the researchers synthesized a number of small bacteria-killing proteins called “antimicrobial peptides.” These proteins are important to antimicrobial and antibiotic research as they may offer new ways in which to treat diseases caused by antibiotic resistant bacteria. However, these molecules are difficult to cultivate as traditional protein synthesis relies so heavily on bacteria. Getting the bacteria to make these proteins is the molecular biology equivalent of installing a self-destruct button stuck on the “on” position. Instead, as cell-free protein expression is abstracted from the requirement of cell growth, the researchers were able to use their cell-free expression system to make large volumes of various antimicrobial proteins, ready for further research.
Cell-free protein expression is an important application in synthetic and protein biology whose potential is yet to be realized. It is apparent that the recent research by Des Soye and colleagues provides a significant optimization in the space, partly realized by Twist Bioscience’s ability to provide sequence-perfect, ready-cloned genes at low prices. It is possible that this work will aid in realizing the potential of synthetic biology and simultaneously enable protein researchers to access new, complex, difficult-to-express proteins that aid in fields like health, antimicrobial research and beyond.
Featured image: Colonies of bacteria in culture, medium plate (Adobe Stock)
What did you think?
Like
Dislike
Love
Surprised
Interesting