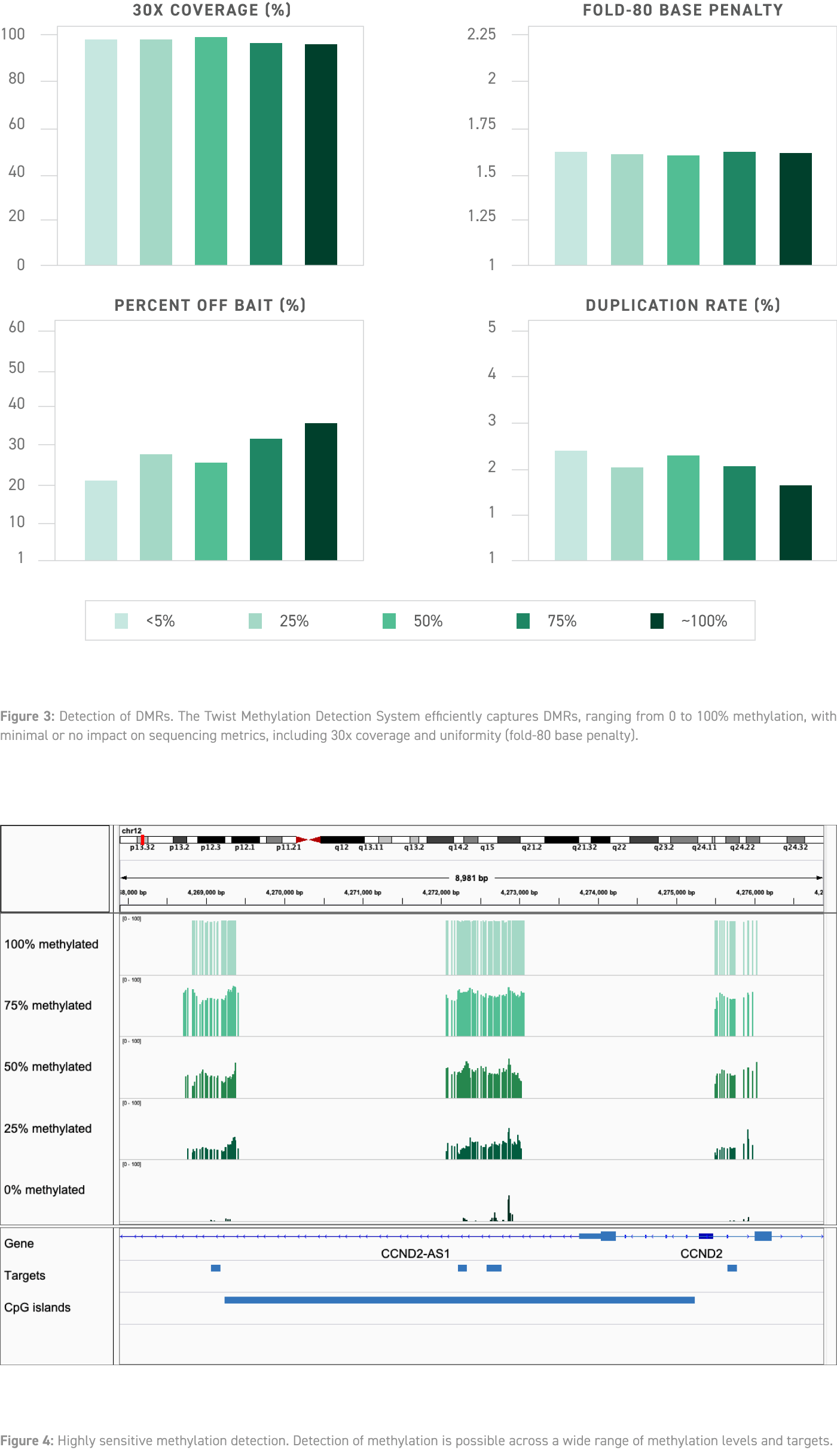
The Twist NGS Methylation Detection System provides a robust, end-to-end sample preparation solution for identifying methylated regions in the human genome. The workflow employs a unique enzymatic process from New England Biolabs® that is much less damaging to DNA, alongside Twist‘s Custom Methylation Panel design.
Whether you are investigating cellular differentiation or screening liquid biopsies for cancer, the system offers the most efficient methylation detection available.