- OVERVIEW
- DATA
- ORDERING & SPECS
- RESOURCES
Protein engineering screens using single site variant libraries allow researchers to explore a protein’s sequence space and investigate the relationship between sequence and protein structure and function. Many methods exist to generate variant libraries, but they generally have significant drawbacks and limitations, including lack of codon control, sequence biases, and incomplete generation of desired variants.
Twist Bioscience Site Saturation Variant Libraries leverage massively parallel oligonucleotide synthesis using a silicon-based DNA synthesis platform and extensive molecular biology expertise to systematically and precisely construct variant libraries. Twist libraries are NGS-verified to confirm that all desired variants are present in the correct ratios.
KEY BENEFITS
Precisely Crafted Variant Libraries
- Complete control over codon usage (all 64 codons available)
- High uniformity of variant representation at every site
- No unwanted codons or premature stop codons
Verified Quality
- Rigorous quality control
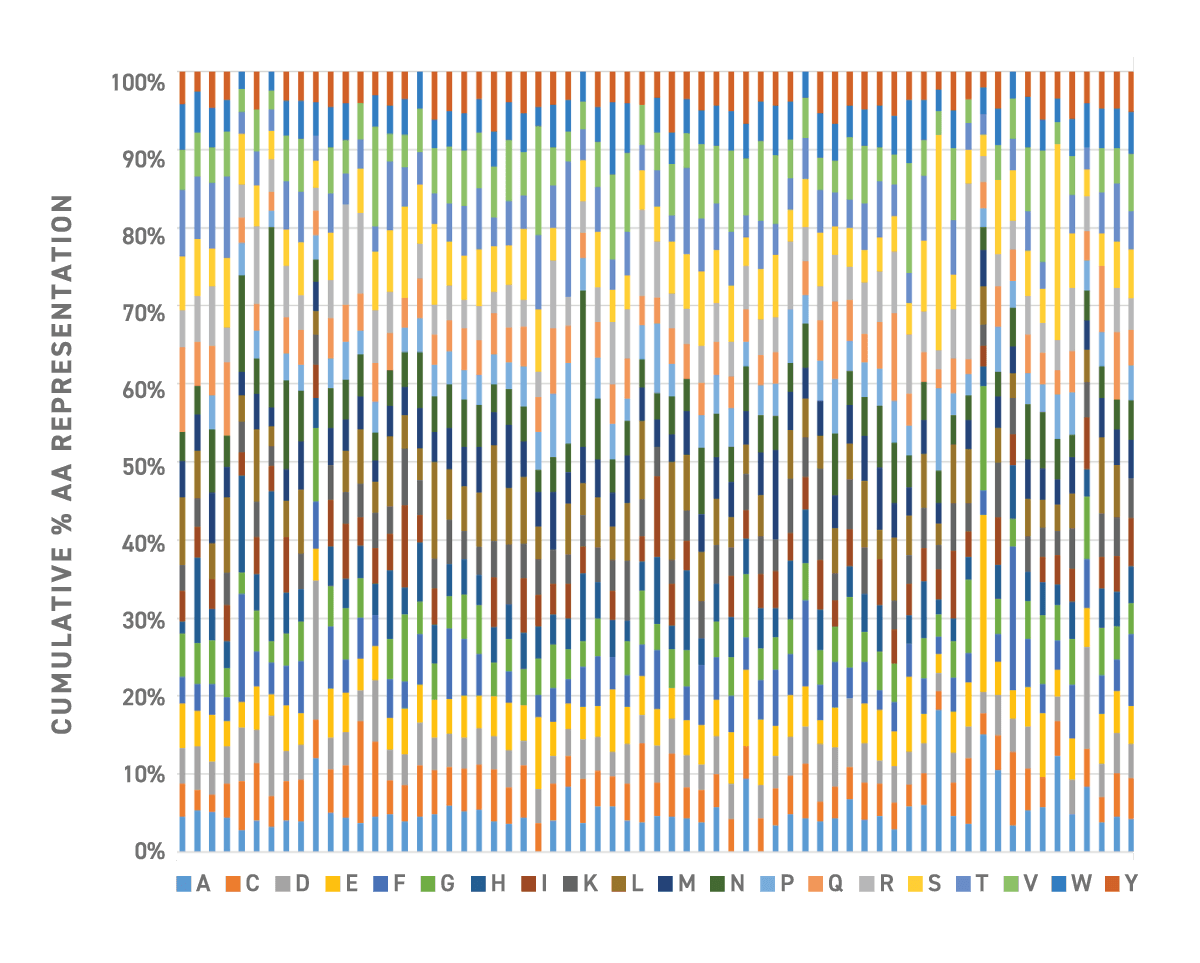
- Verified uniformity of variant representation by NGS analysis of modified region
- Representation of each site normalized by mass
Flexibility
- Pooled library or pooled per position delivery options
- Screen one or up to all 20 different amino acid combinations at each position
- Investigate sequence variants and/or indels
Site Saturation Variant Libraries Submission FormUnleash the full power of a well-designed library. With Twist Bioscience, you are assured that the library you design contains the specific modifications, exactly where you want them, saving you valuable resources for the downstream screening process. Submit your Site Saturation Variant Library designs
Improved Library Generation Fueled by a Silicon-Based DNA Synthesis Platform
The precision of Twist Bioscience’s technology allows for the sampling of large variant space with predefined base by base composition and ratio control. Other library fabrication methods, such as NNN and NNK, result in redundant codons that lead to poor control over diversity and uneven distribution of amino acids.
- Terms & Conditions
- Policy on Unsolicited Submissions of Information
- Privacy Policy
- Regulatory and Quality Information
- Security
- Payment Information
-
© 2025 Twist Bioscience. All rights reserved.