10 Developments Ready to Advance Next Generation Sequencing in 2019
The biological sciences are being transformed by the adoption of next-generation sequencing (NGS). Researchers now perform a wide variety of applications and study biological systems at a level never before possible by taking advantage of NGS’s ultra-high throughput, scalability, and speed. Using innovative sample preparation and data analysis, scientists now have at their disposal more options for applications, including RNA sequencing to examine new RNA variants, rapidly sequencing whole genomes and more precisely targeting genomic regions.
The innovation driven by NGS in 2018 will continue in 2019, offering great promise in a variety of areas. In 2018, researchers began to develop programs to make diagnostic testing more easily available for patients enrolled in the public health plans of the Centers for Medicare and Medicaid Services (CMS), opening up its benefits to potentially hundreds of patients. New genetic data analysis such as that from the Cancer Genome Atlas is revolutionizing the deep analysis of various cancer types and driving the development of precision medicine. Funding from a wide variety of sources will continue to support programs -- to improve whole genome sequencing, to release the first FDA-designated public genetic variants repository and to introduce new processes that drive down the cost of sequencing -- to name just a few remarkable advances in the field.
Read on to discover how these initiatives promise to shape the future of NGS development in 2019:
1. CMS Extends Health Plan Coverage for NGS Testing

DNA model (Source: Adobe Stock)
The reach of NGS diagnostic testing is set to dramatically expand in 2019 as the CMS said it will cover NGS testing for patients with advanced cancer in its health insurance plans.
“We want cancer patients to have enhanced access and expanded coverage when it comes to innovative diagnostics that can help them in new and better ways,” Seema Verma, CMS Administrator, said in a news release. “That is why we are establishing clear pathways to coverage, while at the same time supporting laboratories that currently furnish tests to the people we serve.”
The CMS said these tests, when used as a companion diagnostic to identify patients with certain genetic mutations that may benefit from U.S. Food and Drug Administration-approved treatments, can assist patients and their oncologists in making more informed treatment decisions. Also, when a known cancer mutation cannot be matched to an approved treatment, the results from the diagnostic lab test using NGS can help determine a patient’s candidacy for cancer clinical trials.
2. The First Data Based on the Pan-Cancer Atlas Is Published
Cancer cell (Source: Adobe Stock)
The Cancer Genome Atlas (TCGA) is a collaborative project between the National Cancer Institute and the National Human Genome Research Institute, which after multiple years of research, released the Pan-Cancer Atlas in 2017. In 2018, 25 papers were published together in the journal Cancer Cell, representing the first research published that utilizes the 2.5-petabyte, publicly available database.
The Pan-Cancer Atlas is a comprehensive, multi-dimensional map of aberrations to the genome and epigenome of 33 different types of cancer. In total, data was derived from tumor tissues and normal tissues of over 11,000 patients.
Research highlights include a study that compared 971 adenocarcinomas from different tissues in an effort to understand the factors differentiating gastrointestinal tract adenocarcinomas from other cancer types. Another study showed strong correlative links between whole chromosome or chromosome arm imbalance and somatic mutation rates, and the expression of proliferation genes across the 33 cancer types in the Pan-Cancer Atlas. Importantly, the Pan-Cancer Atlas remains publicly available for further mining, and promises to continue to revolutionize the deep analysis of various cancer types.
3. UK Project Sequences 100,000 Whole Genomes
DNA sequencing (Source: Adobe Stock)
In December, the British government announced it had reached its six-year goal of sequencing 100,000 whole genomes from National Health Service patients. With the project’s completion, this year the UK intends to apply whole genome sequencing at scale in direct healthcare, as well as provide access to high quality, anonymized, or “de-identified” clinical and genomic data for research aimed at improving patient outcomes. The project is intended to support a NHS Genomic Medicine Service, which will provide equitable access to genomic testing to NHS patients starting in 2019.
Genomics England, set up by the NHS, developed 13 NHS Genomic Medicine Centres (GMCs) to support the project, as well as a state-of-the-art sequencing center run by Illumina, and an automated analytics platform to return whole genome analyses to the NHS.
“The results, which will continue to be returned to patients, show how genomic medicine can transform lives, bringing quicker and better diagnoses and increasing the number of patients surviving cancer, and the opportunity now is for the NHS to turn this research into reality by introducing sequencing technology as part of our world-leading NHS Genomic Medicine Service,” said Professor Dame Sue Hill, Chief Scientific Officer for England and Senior Responsible Officer for Genomics at NHS England.
4. NIH Funding for Genome Centers Pushes Precision Medicine Discoveries

Laboratory work (Source: Adobe Stock)
The National Institute of Health (NIH) All of Us research program awarded $28.6 million in September to establish three genome centers which will begin to generate genomic data from biosamples contributed by the program’s participants. This information will become a critical part of the program’s precision medicine research platform, a national resource to support studies on a variety of important health questions.
The centers, led by Baylor College of Medicine, the Broad Institute, and the University of Washington, will generate and analyze genomic data from biosamples contributed by participants in the program.
All of Us is aiming to sign up at least 1 million participants for its precision medicine research platform, including from groups that have been historically underrepresented in research. So far, it has registered more than 110,000 people; 60,000 of whom have completed all elements of the core protocol. Their data will be available to researchers for health-related studies. The centers will produce genomic data for the research effort and will analyze data that will be returned to participants. Participants interested in receiving results will initially obtain information from a list of 59 actionable disease risk genes, as defined by the American College of Medical Genetics and Genomics, as well as pharmacogenomic results. In the future, they will also obtain information about their ancestry and traits.
“Diversity is a hallmark of this effort. We strive for diversity of people and also diversity of data types, so researchers can understand the many factors that influence health and health outcomes for each of us,” Eric Dishman, director of the All of Us Research Program, said in a statement. “Bringing on these new partners is an important milestone for our program as we look to add genotyping and whole genome sequencing data to the many other data types we’re already collecting.”
5. Ancient Offspring of Neanderthal Mother, Denisovan Father Determined
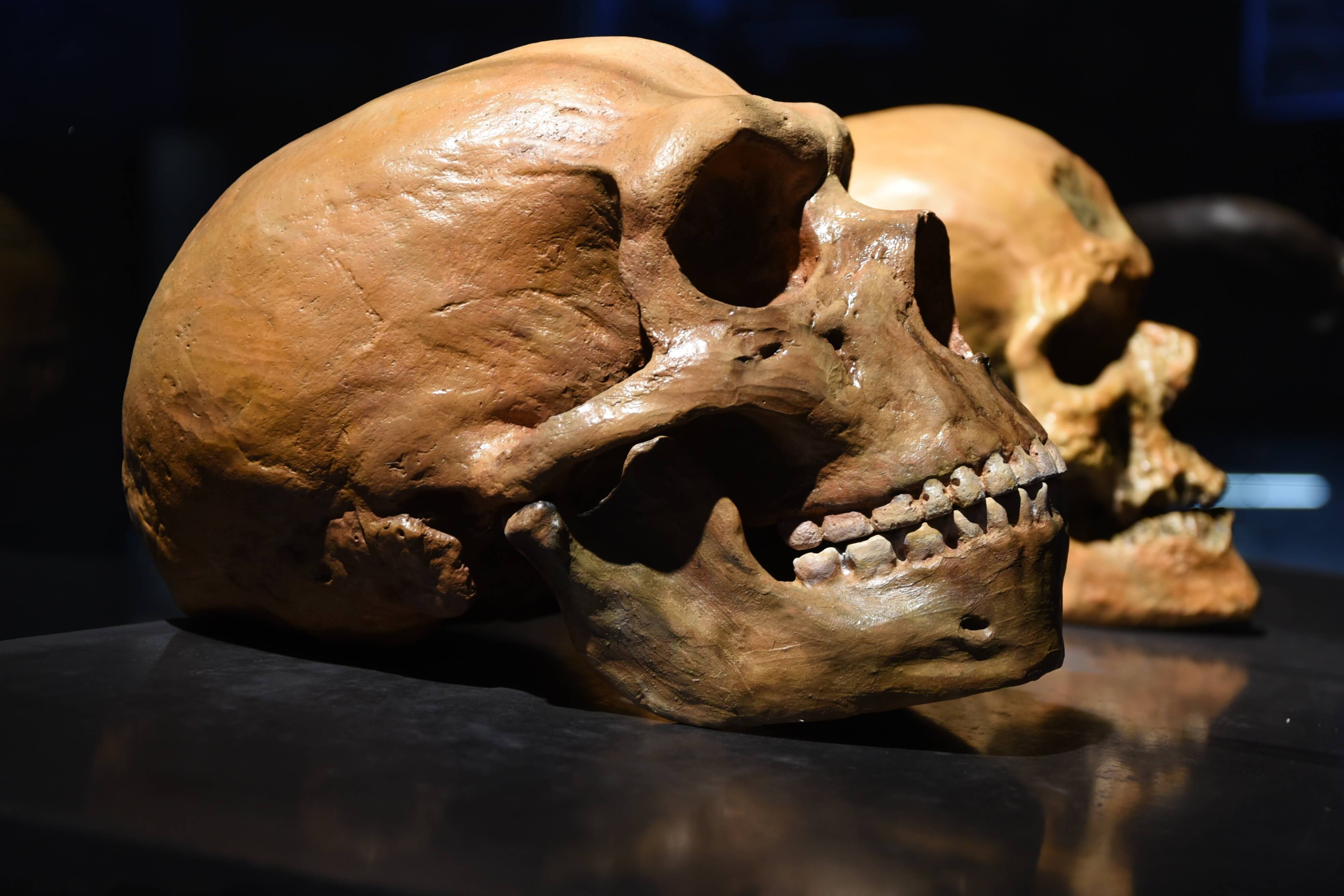

Neanderthal in front of human skull (Source: Adobe Stock)
Researchers of the DNA of our ancient ancestors in 2019 will have greater understanding of the complexities of our evolutionary development based on an investigation suggesting that mixing among archaic and modern hominin groups were likely common when and where the populations overlapped.
Using genome sequencing, researchers were able to determine that a bone fragment from an individual in a cave in Siberia had a Neanderthal mother and a Denisovan father. Neanderthals and Denisovans diverged from each other about 390,000 years ago, although there has been evidence of admixture between the two ancient hominin groups after that split.
Researchers at the Max Planck Institute for Evolutionary Anthropology sequenced the genome from a long bone in Denisova Cave in Siberia that was more than 50,000 years old. In their report in Nature, they showed the individual was a first-generation Neanderthal-Denisovan offspring. After six DNA extractions to make 10 DNA libraries, the researchers sequenced the Denisova 11 genome to a mean 2.6-fold coverage on the Illumina MiSeq or HiSeq 2500 platforms. Based on the coverage of the X chromosome as compared to the autosomes, they concluded Denisova 11 was female. Using three different methods, the researchers estimated that contaminating present-day human DNA made up 1.7 percent of their data at most.
"We knew from previous studies that Neanderthals and Denisovans must have occasionally had children together," co-first author and Max Planck researcher Viviane Slon said in a statement. "But I never thought we would be so lucky as to find an actual offspring of the two groups."
6. FDA Releases Guidelines for Development of NGS-Based Tests

DNA model (Source: Adobe Stock)
With the rapidly decreasing costs of NGS technologies, genetic screening is becoming an increasingly commonplace aspect of patient treatment. This trend promises to accelerate in 2019 as the U.S. Food and Drug Administration (FDA) released a set of guidelines in April for the design, development and validation of tests that utilize NGS and the use of publicly available databases of genetic variants to support the soundness of data derived from the tests.
The two key guidelines outline the FDA’s expectations when assessing whether pre-market applications are safe, accurate and reliable for the detection of specific genetic variants in a clinical setting. “The new policies issued today provide a modern and flexible framework to generate data needed to support the FDA’s review of NGS-based tests, and give developers new tools to support the efficient development and validation of these technologies,” said Scott Gottlieb, FDA commissioner.
“Information about genetic variants is generally stored in a manner that is not publicly accessible,” Jeff Shuren, director of FDA's Center for Devices and Radiological Health, said in a statement. “Today’s release will help change this paradigm by encouraging data sharing and the accumulation in public databases of evidence supporting the clinical validity of genomic tests to help provide an even more efficient path to market."
7. African Pan-Genome Analysis Reveals that Reference Genome Sequences may not be Adequate
African continent illustration (Source: Adobe Stock)
The human reference genome is a digital database representative of the complete set of genetic information belonging to humans. Data from the deep sequencing of a small number of individuals' DNA compromises the database. However, the best utilization of sequencing data for population genetics, healthcare, and medical screening relies on a reference genome that is representative of all populations.
A team at John Hopkins University analyzed sequencing data at between 30x and 40x depth from 910 individuals of African descent, representing 1.2 trillion paired end reads. Analyses revealed almost 300 million bases in 125,715 contigs did not align to the most recent reference genome release (GRCh38). This bares a significant result for the future of human genomics, as a single reference genome may not be adequate for all demographics. “Instead, a better approach may be to create reference genomes for all distinct human populations, which over time will eventually yield a comprehensive pan-genome capturing all of the DNA present in humans” said Steven Salzberg, co-author of the analysis.
As only a small number of individuals are used to create GRCh38, some pan-genome contigs have been lost. The African pan-genome containing the new contigs identified by the team at John Hopkins University represents a reference genome that contains 10 percent more genetic information than the current generation human reference genome.
8. France’s Personalized Medicine Project to Begin Sequencing Genomes

Modern laboratory (Source: Adobe Stock)
France’s national personalized medicine initiative will start sequencing patient samples in the first quarter of 2019 with two pilot genomic platforms, a spokesman said in December. The project, called the Plan France Médecine Génomique 2025, will use Illumina NovaSeq instruments on both platforms, with each expected to generate roughly 18,000 genomes from patient samples by 2021.
“All data will be sent to a central repository in a single place, and this will become a public resource for both researchers and private partnerships,” said Jean-Yves Blay, director of Centre Léon Bérard, a comprehensive cancer center based in Lyon in Southern France. "France has had whole-genome sequencing installed for the past decade, but following this plan, we will have easier access to this infrastructure to help patients."
Blay is also the scientific manager for Auragen, one of the two pilot platforms that will begin operations next year. The sequencing platform is housed and administered by the genetic service of Hospices Civils de Lyon. The other platform is the Sequencing Omics Information Analysis (SeqOIA) cooperative health group, based in Evry, a suburb of Paris.
9. FDA Recognizes ClinGen Database as Public Genetic Variants Repository

DNA molecules with digital data background (Source: Adobe Stock)
The FDA announced in December that it is recognizing the Clinical Genome Resource Consortium's ClinGen database as the first FDA-designated public genetic variants repository. The recognition of the database, which contains information about genes, genetic variants and their relationship to disease, aims to encourage the development of novel diagnostic technologies that scan a person’s DNA to diagnose genetic diseases and guide medical treatments.
The approval is the first of its kind issued through agency's Human Variant Database Program, which is intended to help reduce regulatory burdens on companies developing NGS-based genetic tests and foster the advancement of precision medicine.
The agency's recognition "highlights for the genetics community our goal of developing authoritative sources of clinical genetics that is publicly available to them," Sharon Plon, ClinGen principal investigator and Baylor College of Medicine geneticist, said in a statement. "ClinGen is filling a critical need in the substantial challenge of gathering data and connecting it with the community for use in supporting genomic medicine and research."
10. Illumina Broadens Access to Long-Range Sequencing with $1.2 Billion Purchase of Pacific Bioscience
Computer and Genome DNA Sequencer (Source: Adobe Stock)
In a move to accommodate the growing whole genome sequencing market, Illumina announced in November plans to buy rival Pacific Biosciences for approximately $1.2 billion. Illumina said that PacBio's long-read sequencing technology will complement its own short-read sequencing platforms and will allow it to provide integrated workflows and innovations that bring together the best of both technologies.
Illumina CEO and President Francis deSouza touted PacBio's accuracy, technology roadmap, and potential clinical applications as key to why the company wanted to acquire it. In addition, deSouza said the acquisition would broaden Illumina's market into areas such as de novo assembly of plant and animal genomes, functional genomics, tissue transplantation, and pharmacogenomics — applications that he said require "uniform, unbiased coverage in highly repetitive regions, which long reads are best suited to provide."
DeSouza added that the long-read sequencing market had a size of around $600 million in 2017 and is expected to grow to $2.5 billion in 2022.
With the rapid pace of change brought on by NGS in 2018, we can expect to see the momentum continue in 2019 in research, testing and clinical practices for a myriad of applications and benefits.
Featured image: DNA sequence (Source: Adobe Stock)
Check out Twist on Twitter, Linkedin, Facebook and YouTube.
What did you think?
Like
Dislike
Love
Surprised
Interesting