Mapping Traits to Genomes Made Simple with CRISPR Tools and Twist Bioscience Oligo Pools

Mapping exactly which parts of the genome control diseases and disorders is a complex task, especially considering the majority of such conditions are impacted by multiple genetic elements simultaneously. Traditional methods often have poor resolution, and can take years. In a recent study published in the journal Cell, Elion Sharon, PhD, and colleagues at Stanford University used oligo pools synthesized by Twist Bioscience to develop a proof of principal for a new method of genetic mapping based on the CRISPR gene editing tool. Twist Bioscience oligo pools provided the researchers the accuracy to map traits to the genome at single base resolution.
Our genomes are the foundation of who we are. Encoded in DNA, packaged into 23 chromosome pairs, and locked away in the nuclei of our 37 trillion-some cells, is the information that defines our “traits.” Traits are personal characteristics that are genetically traceable, like eye color. Uncovering what parts of our genomes encode such traits and how they’re inherited from our parents is an essential scientific pursuit. In the same way, genetic diseases are also traits. Fully understanding the genetics underlying such diseases is pivotal to our ability to anticipate, treat and manage them.
Genetic traits can be bundled into categories. “Mendelian traits” are typically controlled by a single gene and manifest as discrete categories. A good example is your blood type. As genetic research has improved, traits that were previously believed to be Mendelian, like hair color or the ability to roll one’s tongue, have been found to in fact be “polygenic.” Polygenic traits are inherited and controlled through a cluster of genomic factors and the variation of one or more of these factors has varying effects on the trait’s manifestation.
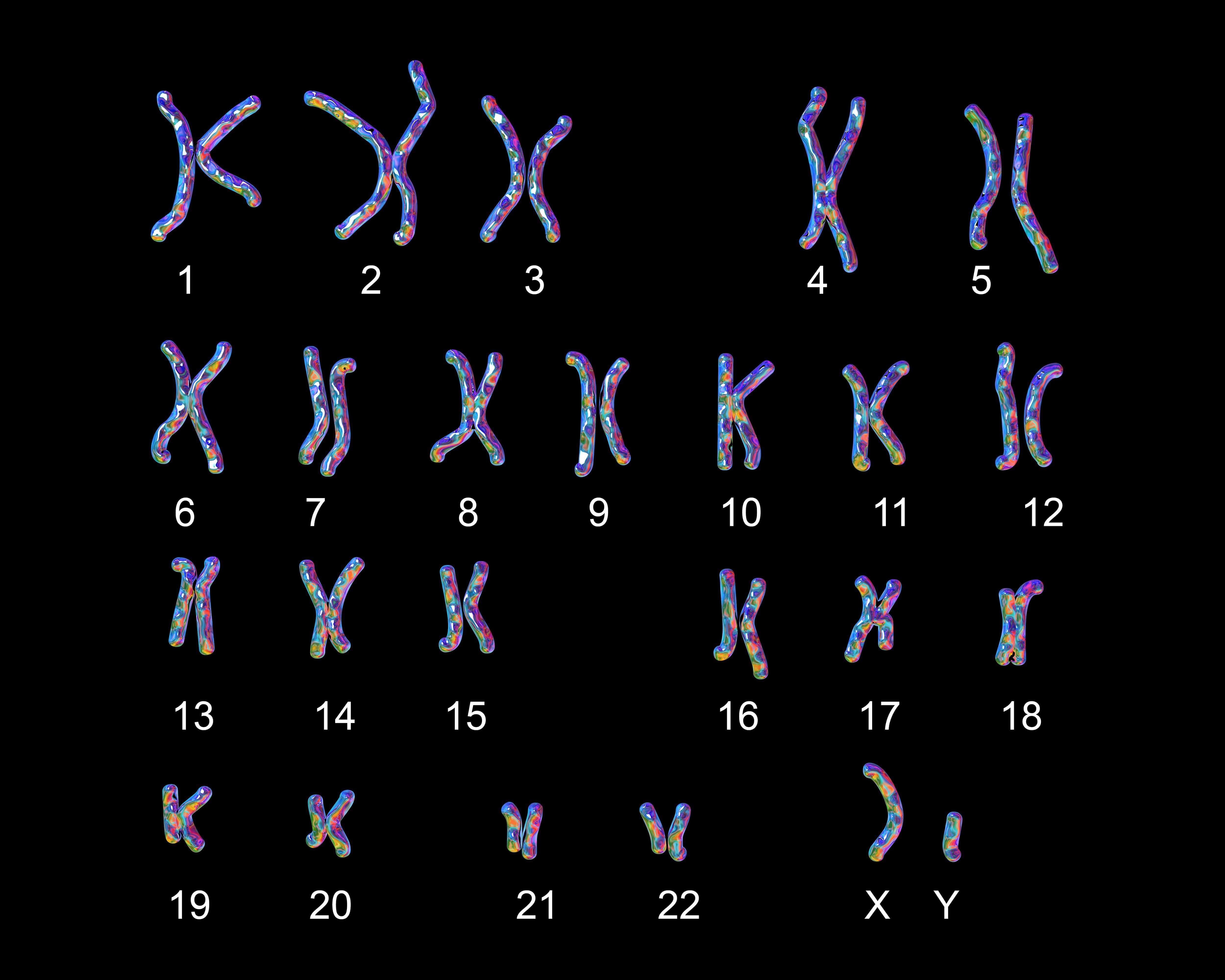
Together, your 23 chromosomes contain almost 6.5 billion base pairs of DNA. Diagnosing exactly how traits are inherited, by understanding which regions control a particular trait, is a challenge.
Most diseases and disorders that have a genetic component are polygenic, and their severity and accompanying complications exist on a bell curve. A good illustration of this is Type 2 diabetes, where individuals show highly variable symptoms and disease progression. Such genetic complexity means that identifying and understanding the full gamut of genetic elements controlling a given polygenic trait is a considerable challenge.
A region of the genome that can be experimentally associated with a trait is called a quantitative trait locus (QTL). Traditionally, cross-breeding experiments in model laboratory organisms would provide an understanding of trait inheritance. QTLs can then be discovered by comparing regions of difference between the genomes of organisms with inherited traits with statistical analyses of variance. Similarly, QTLs can be discovered in humans by using familial genetics and homology, where similarity and comparability between the genomes of relatives, or related species, allows for cross-referencing.
However, QTL mapping in this manner is inefficient, with only small numbers of maps pinpointing actual variants that cause disease (instead maps will identify potential target regions). Mapping often takes years of effort, especially for slow breeding species. Additionally, the amount of genomic elements controlling a given trait is rarely known, so depth and completeness of understanding is tricky to gauge.
In a recent study published in the journal Cell by Elion Sharon and colleagues at Stanford University, new methods are outlined that could dramatically accelerate and simplify the process of QTL mapping. Their approach uses the precise genome editing tool, CRISPR to make genome edits in single cells in high throughput, utilizing oligonucleotides synthesized by Twist Bioscience.
When CRISPR makes a double stranded cut in the genome, the resultant DNA damage is repaired by the cell. This repair process can be used to integrate new genetic information into the cut site. The most precise version of this technique uses a repair process known as Homology Directed Repair (HDR), in which integrated sequences contain homology to the DNA sequences on either side of the CRISPR cut site. By performing HDR in high throughput, every variant from two-parent cells can be integrated into a cell population one by one, and the effects of these variants on a given trait at a cellular level scored.
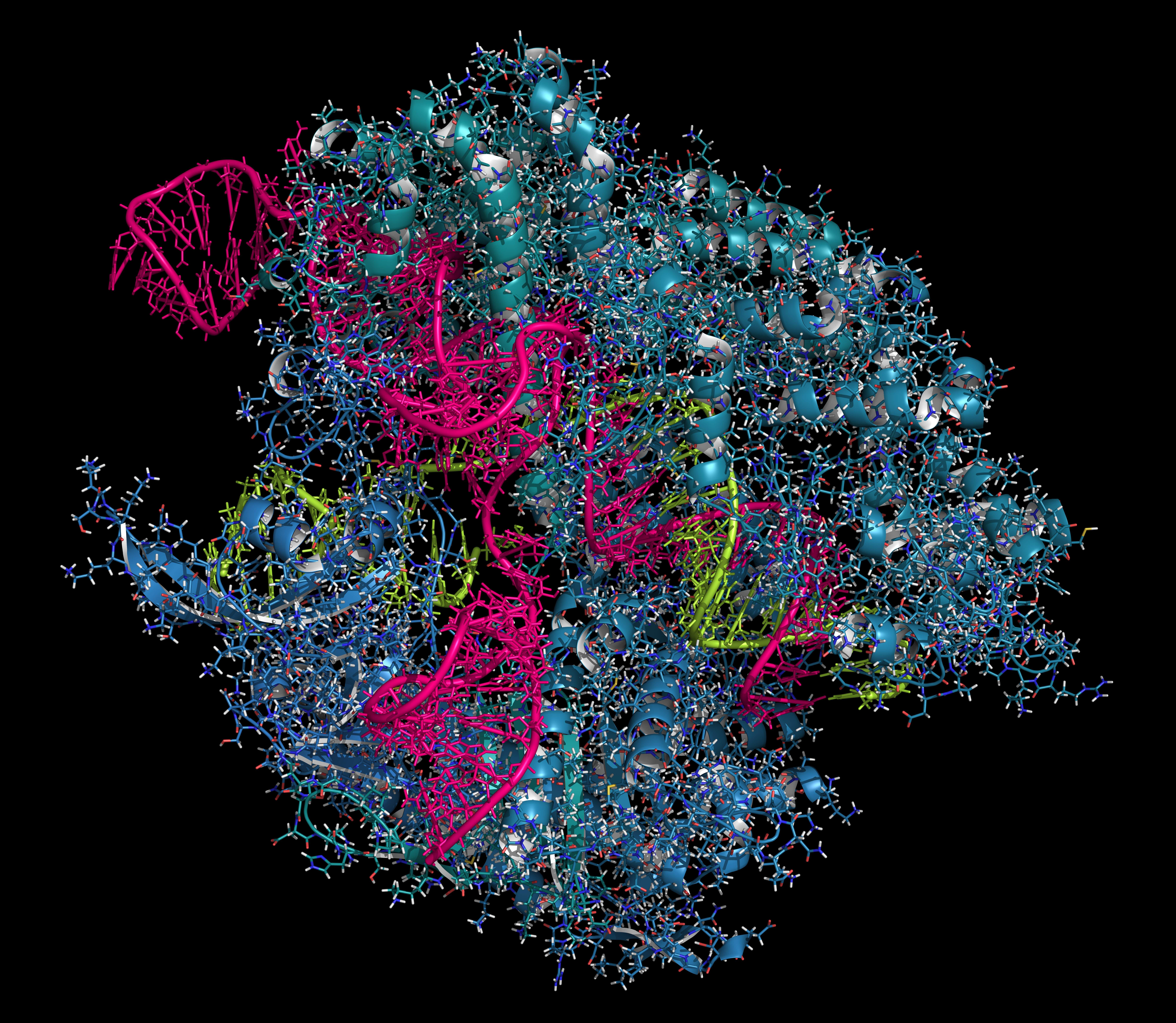
CRISPR is a highly precise gene editing tool that uses an RNA molecule to guide it to a specific point in the genome.
However, in order to achieve this in high throughput, the researchers had to circumvent a crippling inefficiency issue with CRISPR. To achieve HDR, the molecular scissors of CRISPR (an enzyme called Cas9) is transfected into the cell alongside its guide RNA, and a corresponding template for HDR. For a successful experiment, a corresponding guide RNA and HDR template need to end up in the same cell at the same time. In high-throughput experiments, tens of thousands of guides and inserts are introduced into a population of cells simultaneously. The chance that a gRNA and a HDR template both target the same region of the genome and make it into the same cell at the same time are extremely small.
Instead, the researchers designed a tool, called “CRISPEY,” that links a guide RNA with its corresponding HDR template so the two requirements always integrate into the same cell. This was achieved with a tool derived from viruses - retrons. Retrons are RNA elements that are reverse transcribed to DNA in high abundance. As CRISPR already requires a guide RNA to direct its cut site, the researchers extended the sequence of a guide RNA beyond its Cas9 interacting region, to include a retron. When the retron is reverse transcribed, it produces the DNA template for HDR in high abundance.
Therefore, in theory if a cell already makes an enzyme for reverse transcription, only the Cas9 enzyme and its guideRNA-HDR template fusion need be transfected into the cell. Once in the nucleus, the template DNA is then made from the retron, the Cas9-guide RNA cuts the target site as intended, and the cell’s own DNA repair machinery integrates the retron-encoded DNA into the targeted site.
To provide a proof of principle that their method works, the researchers utilized Twist Bioscience oligonucleotide synthesis to synthesize a pool of 32,000 guide-HDR template fusions, targeting a yeast genome. In such CRISPR-based QTL mapping experiments, it is essential that the guides and templates are accurate, as, at the genome level, single nucleotide differences can manifest dramatic changes in a given trait. Therefore, if the oligonucleotides encoding the retron contain errors, it is difficult to distinguish whether a genome-trait relationship observed is a true result or a false-positive. Twist Bioscience provides oligo pools with industry leading accuracy, allowing researchers to ensure that their observations provide true results.
When testing their methods in yeast, the researchers targeted approximately 16,000 variants to better understand the underlying genetic elements that control a yeast cell’s fitness (its ability to outcompete other yeast cells for resources). They found that their method integrated new genetic information with 96% efficiency, showing that retron and CRISPR-based HDR is a powerful tool for QTL mapping, especially due to its scalability. 171 variants were observed to harbor an increase of >1% fitness, and 17 harbored fitness increases of >5%. More importantly, the genetic regions controlling these significant increases in fitness could be identified at the single base-pair level, providing exceptional resolution.
Next, the researchers aim to transfer their methods over to mammalian cells. As already described, accurate oligonucleotides encoding guide RNA-HDR template fusions will be essential to successful high throughput QTL mapping. Twist Bioscience provides accurate oligonucleotides at any scale, making them highly transferable to such experiments. In the future, advances in QTL mapping will likely play an important role in effective personalized medicine. By precisely mapping genomic effectors of polygenic diseases, better detail about how a patient’s disease manifests, progresses, and responds to treatment can be gained.
Ultimately, base-resolution QTL data will compliment next-generation sequencing. Regions of a patient’s genome can be pinpointed for sequencing to identify their specific trait genotype at far lower costs than whole genome sequencing. Such personalized medicine will allow for optimal treatments, better intervention, and better prognoses for countless individuals.
Featured image: Insertion of new genetic information concept (Adobe Stock)
What did you think?
Like
Dislike
Love
Surprised
Interesting