Opening New Horizons, Scientists Achieve Complex Logical Processing with Biological Computers

Researchers from Université de Montpellier in France have developed and tested more than a dozen cell-based biological devices that can work together to process complex logic. This work, recently published in Nature Communications, increases the functionality of biological computers and expands their abilities as synthetic tools to support applications in fields such as bioprocessing, healthcare and material science. To construct the biological devices, which depend on high reliability, the researchers used Twist Bioscience high-quality gene fragments. Logical processing within the devices depend on the presence of very specific DNA sequences, and therefore required high-quality DNA for construction.
Synthetic biology uses natural biological parts to construct unnatural biological tools. In recent history, the computer is the tool that has impacted society perhaps more than any other, and synthetic biologists took notice. Among their many attributes, computers are useful because of their ability to quickly calculate and operate using complex logic, and control mechanical devices with an array of functions. While computers can only receive inputs tied to electricity, biological computers accept biological molecules as inputs and can make new ones as outputs. These biological computers could function as diagnostics, drug delivery devices, sensors, and cell classifiers, among many other uses.
One of the critical components of any computer is an ability to perform logical operations. In fact, even basic machinery, like a toaster, relies on simple logical processing. For example: IF the toaster has been toasting for a set amount of time, THEN turn off. However, to achieve more sophisticated functionality, biological computers and other types of biomachines need to be able to perform complex logic by processing combinations of AND, OR, and NOT operations (also known as boolean operations, which combined are boolean functions).
Over the last few decades, synthetic biologists have constructed biological computers that can process logical operations with higher and higher complexity enabling impressive functions like the ability to play Tic-Tac-Toe, identify cancer cells, induce cell death, and deliver drugs. Each biological computer is composed of cells that are encoded with a logic gate, which under certain conditions (like the presence of certain mRNAs) will output a response (like the transcription of a gene). Thus far, logic gates within biological computers can reach a fair amount of complexity, however, each one is ‘made-to-order.’ In other words, if even a slight change to the logic gate within a biological computer needs to be made, an entirely new one needs to be constructed. Modularity of cell-based logic gates would allow for quick changes in logical processing and increase the problem-solving capabilities of biological computers overall.
Similar to how paint can be mixed into any color using only primary colors, any logical function can be processed using a combination of a finite set of logic gates. Researchers from Université de Montpellier used this concept to develop a set of cellular logic gates that can be used in thousands of combinations to process complex logic. Their work was recently published in the journal Nature Communications, in a paper entitled, “Hierarchical composition of reliable recombinase logic devices.” The researchers scaled their system to accept four inputs, which translated into a minimum of fourteen distinct logic gates that in different combinations (a total of 16,384 combos) could process any 4-input logic function. The set of cellular logic gates are an ‘off-the-shelf’ type of solution for the fast construction of biological computers with abilities to process complex logic and can be scaled further to achieve higher level logic.
The team decided to develop recombinase-based logic gates, which is a tried and true method to achieve reliable processing of complex logic. Recombinases are natural enzymes found in bacteria, eukaryotes, and viruses that initiate DNA recombination (rearrangement of DNA by exchange of DNA strands) at specific sequences. Depending on the placement of the recombinase recognition sequences, addition of a recombinase can result in insertion, excision, inversion, or translocation of a piece of DNA.
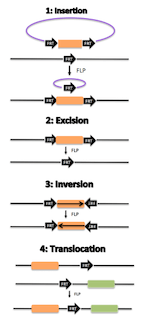
Example of FLP recombinase functionality. At FLP recognition targets (FRTs) FLP induces DNA recombination. Depending on the placement and direction of the FRTs, the insertion, excision, inversion or translocation of DNA is possible. (Source: Wikimedia Commons)
DNA excision occurs when two recombinase recognition sites flank that piece of DNA. The researchers relied on this property of recombinases to construct logic gates in bacteria. If gene expression is the output of the logic gate, recombinases can be used to cut out DNA sequences that start or block transcription. To block transcription in the presence of a recombinase, the recombinase recognition sites (RRS) would be placed flanking the gene’s promoter sequence, which is needed to start transcription (example DNA strand: ---RRS-Promoter-RRS-Gene---). To start transcription, recombinase recognition sites would flank a terminator, which are sequences of DNA that stop transcription, after the promoter (example DNA strand: ---Promoter-RRS-terminator-RRS-Gene---).
Within bacteria there are many unique recombinases that recognize distinct DNA sequences. To construct logic gates that could accommodate four inputs, four distinct recombinases with unique DNA recognition sequences are needed. With thoughtful placement of recombinase recognition sites, terminators, and promoters, the researchers were able to construct DNA based logic gates that work in combinations to process any 4-input logic function. Their output in this system was the expression of green fluorescent protein (GFP), which they measured with a flow cytometer to verify that the system works and is reliable.
Another advantageous concept of the system that the researchers built, is that the presence or absence of recombinases can be tied to the presence or absence of other molecules. For example, it’s possible to add another DNA fragment to the system that will express a recombinase only if a specific mRNA is present. This means that the inputs to the logic gates can be easily changed while preserving the logic processing capabilities to add even more power to these biological computers.
With biological computers that can be easily adapted to process a vast array of logic functions and accept diverse inputs, synthetic biologists will be able to solve a wide variety of complex problems. While we are still at the beginning of biological computer development, the field is making fast strides towards practical application with the ability to disrupt and revolutionize many fields. In healthcare, diagnostic machines, which detect biological molecules, could be replaced with biological computers. Drug delivery devices could be integrated as combined diagnostics and drug production machines with biological computers. With the ability to control cell behavior, tissue engineering could use biological computers to ‘build’ tissues and organs. The possibilities are endless.
Featured image: DNA sequence / Researcher working with analysis DNA software on laptop
(Source: Adobe Stock)
What did you think?
Like
Dislike
Love
Surprised
Interesting