Rather than a neat, linear sequence of As, Gs, Cs, and Ts, our genome better resembles a bowl of spaghetti: a mess of intermingling loops of DNA strewn about. Or at least that’s the way we viewed it before the advent of fluorescence in situ hybridization (FISH), a method that involves labeling genomic DNA with fluorescently tagged oligos.
Just as cells are not simply bags of enzymes, nuclei are not bowls of chromosomal spaghetti. The genome has a nonrandom three-dimensional organization that determines how and when genes are expressed.
Tools like FISH enable researchers to pinpoint the location of specific DNA sequences in three-dimensional space with ever-increasing detail, so much so that researchers may now study the entire genome with nanoscale resolution. With these genomic pictures in hand, researchers can begin to answer pressing questions that DNA sequencing alone cannot, such as how chromosomal evolution differs across species and how chromosomal architecture influences gene regulation in development and disease.
The rise of spatial genomics has been driven by advances in both imaging technology and synthetic biology. Specifically, advances in the large-scale synthesis of DNA has been catalytic, enabling studies to expand in focus from tens of loci to thousands.
What is Spatial Genomics?
Spatial genomics is the study of the three-dimensional structure of the genome. It spans the hierarchical organization of chromosomes into loops, loops into topologically associated domains, then into compartments, and compartments into territories. But where current indirect methods (i.e., chromatin capture) offer averaged snapshots of these structures, direct visualization methods like FISH can resolve them in single cells, enabling an analysis of how these structures differ between cell types and even cell states. Doing so at genome-scale could reveal how a local change — be it normal genetic variation, a chromosomal rearrangement in cancer, or a virus inserting its genome into a host genome — affects the global genome structure and function.
Spatial genomics can reveal global genome structure and function
A recent example of this is the discovery that genetic variants associated with type 1 diabetes could cause chromatin to misfold, bringing disease-associated enhancers and promoters together into “hyperconnected cliques” and causing aberrant expression of disease-related genes1. Although the researchers initially used chromatin capture to discover these genomic structures, the pictures “painted” by FISH provided explicit confirmation about the role of genomic structure in aberrant gene expression and disease phenotypes.
A deeper understanding of genomic structure-function relationships may also inform synthetic biology. If an endogenous gene’s expression is controlled by its spatial positioning — whether the local chromatin environment is open or closed or which distal elements are looped nearby — so too is the expression of synthetic genes. Advances in the scale of spatial genomics can help guide the rational design of ideal synthetic gene integration sites, and lead to the construction of cell lines with more stable or predictable expression of exogenous genes and pathways.
Oligopaints drive the high-throughput revolution
At its most basic level, the DNA FISH procedure involves hybridizing fluorescently labeled DNA fragments to sample DNA for direct visualization using a fluorescence microscope. As such, the number of fluorescent channels available for fluorescence microscopy — 4 or 5 — has bottlenecked initial multicolor iterations of the technology.
Scientists have developed two primary approaches to overcome this limit. The conceptually simpler approach is sequential imaging, in which the same sample is hybridized and imaged in multiple successive rounds. This approach scales linearly, resolving only 24 (4*6) unique targets in a single experiment with four color channels and six imaging rounds.
Going beyond dozens of targets to hundreds or thousands requires combinatorial barcoding. A single target will be probed multiple successive times, each time with a new fluorescent label. Researchers can then target multiple loci with one color. It is impossible to differentiate one gene from another in a single image. However, the unique series of colors that hybridize to the target over multiple imaging rounds come together to form an identifying barcode for each locus.
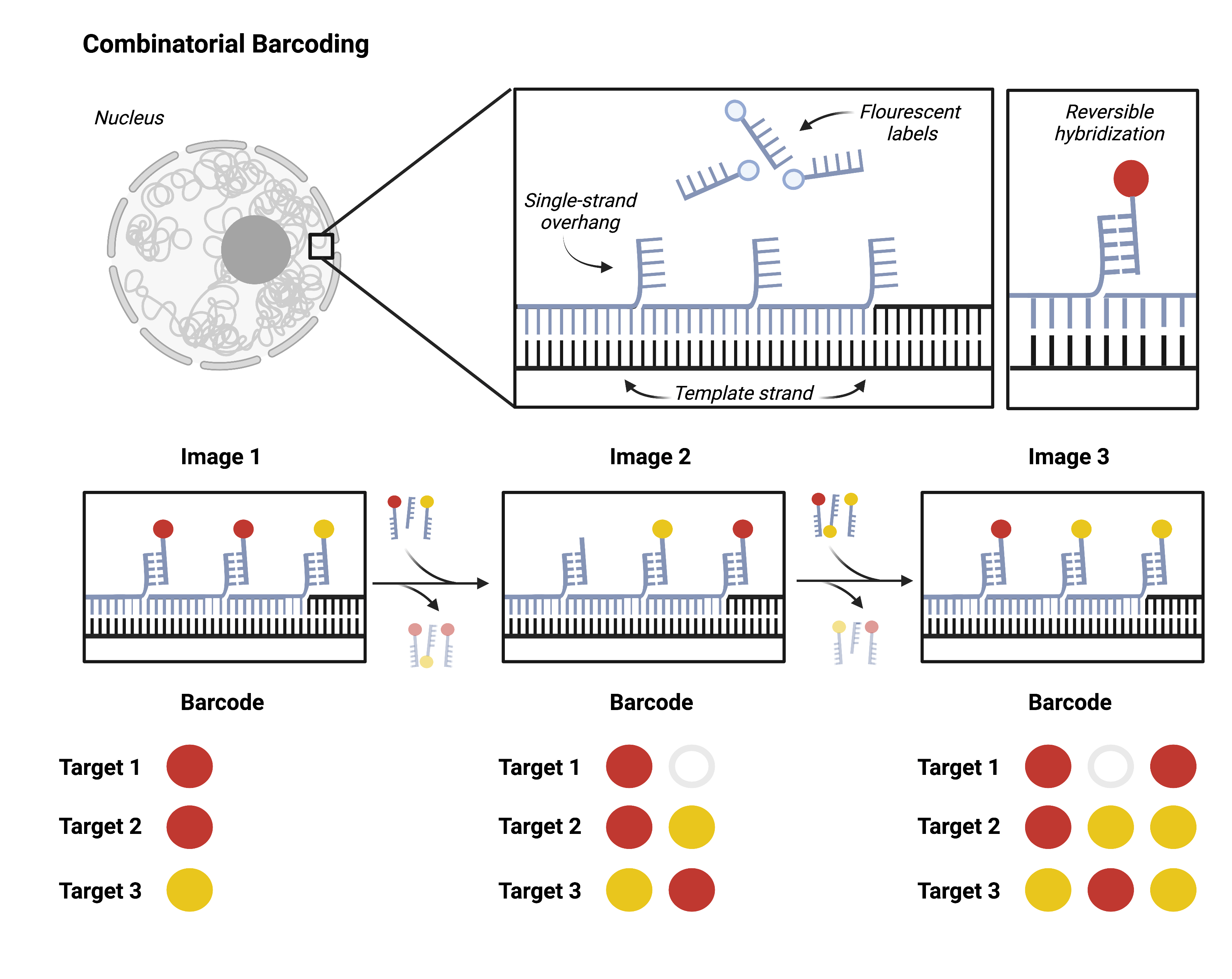
Researchers can design barcodes for each target using combinatorial barcoding and synthesize hybridization probes accordingly. This approach scales exponentially: a single experiment can resolve 256 (28) unique targets with just two color channels and eight imaging rounds. DNA MERFISH and DNA seqFISH+ are both recent applications of this approach.
A challenge of combinatorial approaches like DNA MERFISH and DNA seqFISH+ is cost, which increases as the number of targets, and therefore the number of unique readout probes, increases. New in situ sequencing approaches overcome the cost of scaling by co-opting sequencing-by-synthesis, sequencing-by-ligation, and sequencing-by-hybridization methods to read spatial information from barcoded probes. Part of a suite of spatial genomics tools called OligoFISSEQ (Oligonucleotide Fluorescent In Situ Sequencing), these strategies use the same universal pool of labeled probes to read genomic target locations no matter how many targets are visualized.
Painting with synthetic DNA
DNA MERFISH, DNA seqFISH+, and OligoFISSEQ were all made possible by advances in synthetic DNA — specifically, array-synthesized DNA.
Before the advent of massively parallel DNA synthesis, FISH probes were generated as column-synthesized DNA, by random PCR amplification, or by fragmentation of clone-based DNA (genomic DNA, BAC vectors, and the like). The latter two methods are error-prone and limited by repetitive sequences, resulting in a prohibitively high off-target rate that prevented their application to genome-wide imaging.
Column-based DNA synthesis offered researchers the ability to target the genome precisely with shorter, more effective probes. Short column-synthesized probes also tended to hug their genomic targets better than the longer clone-based probes, reducing inter-probe networking and enabling imaging at superresolution. Yet, the cost of column-synthesized probes prevented scaling. It wasn’t until microarray manufacturers generated the first FISH probe libraries in 2011 that performance and scale came with an accessible price tag.
DNA synthesis technology has evolved significantly over the past decade. On Twist Bioscience’s silicon-based DNA synthesis platform, DNA is synthesized at scales of up to 1 million oligos per chip and lengths of up to 300 bases per oligo. Combinatorial barcoding and in situ sequencing methods require tens to thousands of ~150 bp oligo probes per genomic target to provide the coverage and resolution needed for high-resolution spatial genomics.
Probe synthesis precision and uniformity are also critical factors in experimental success. Precision ensures probes bind specifically to their targets and enables selective targeting of splice variants or single nucleotide polymorphisms. Uniformity guarantees even representation across probes, minimizing probe waste and reducing the likelihood of incompletely resolved targets occurring.
Twist Oligo Pools meet all of these criteria — scale, length, precision, and uniformity — which is why the developers of various functional genomics applications have turned to Twist for their DNA synthesis needs2-5.
If you are interested in using Twist Oligo Pools for your spatial genomics experiment, read more about them on our Oligo Pools for High-Throughput Screens page.
References
- Fasolino, Maria et al. “Genetic Variation in Type 1 Diabetes Reconfigures the 3D Chromatin Organization of T Cells and Alters Gene Expression.” Immunity vol. 52,2 (2020): 257-274.e11. doi:10.1016/j.immuni.2020.01.003
- Takei, Yodai, et al. “Integrated Spatial Genomics Reveals Global Architecture of Single Nuclei.” Nature, vol. 590, no. 7845, 1 Feb. 2021, pp. 344–350, www.nature.com/articles/s41586-020-03126-2, 10.1038/s41586-020-03126-2.
- Su, Jun-Han, et al. “Genome-Scale Imaging of the 3D Organization and Transcriptional Activity of Chromatin.” Cell, vol. 182, no. 6, 17 Sept. 2020, pp. 1641-1659.e26, www.sciencedirect.com/science/article/abs/pii/S0092867420309405, 10.1016/j.cell.2020.07.032.
- Beliveau, Brian J., et al. “Visualizing Genomes with Oligopaint FISH Probes.” Current Protocols in Molecular Biology, vol. 105, no. 1, Jan. 2014, 10.1002/0471142727.mb1423s105.
- Nguyen, Huy Q., et al. “3D Mapping and Accelerated Super-Resolution Imaging of the Human Genome Using in Situ Sequencing.” Nature Methods, vol. 17, no. 8, 27 July 2020, pp. 822–832, 10.1038/s41592-020-0890-0.
- Terms & Conditions
- Policy on Unsolicited Submissions of Information
- Privacy Policy
- Regulatory and Quality Information
- Security
- Payment Information
-
© 2025 Twist Bioscience. All rights reserved.