Detecting the Unexpected

When looking for an answer, something to explain the underlying cause of a disease, researchers often turn to the genome. With more than 3.2 billion bases in the human genome, the first and most financially viable step is to narrow the search to portions most likely to be informative. To do this, researchers can extract specific portions of the genome for sequencing with target enrichment panels. Picking the right panel is key to collecting quality, informative data.
Researchers have two options at this stage: small and targeted panels or whole exome panels. Whereas panels of “clinically relevant” targets offer a narrow and cost-effective view of specific genes, Whole Exome Sequencing (WES) provides a dramatically increased perspective of the genetic landscape. With this broader view, researchers can parse through the data for known pathogenic variants, or piece together patterns that allude to novel gene-phenotype discoveries. Therein lies the power of WES: It provides a breadth of data that can be used for a myriad of applications.
Yet, the value of that data is not a given. Many commercial WES solutions are available on the market, each varying in their quality and content. An ideal WES panel would provide comprehensive, uniform, and on-target coverage of protein-coding sequences. In a word, the ideal WES panel is efficient.
The Twist Exome 2.0 panel is the newest addition to the Twist suite of sequencing tools, and is built to bring a combination of broad coverage and high efficiency to the translational research market.
What is Twist Exome 2.0?
Twist Exome 2.0 is a research use only (RUO) human exome panel that enables targeted sequencing of the whole exome as well as several protein-coding and noncoding regions of the genome.
By their nature, most commercially available exome panels cover nearly all gene-coding sequences in the genome. But, one key differentiating factor between exome panels is how deeply they cover specific genes. Some whole-exome panels focus on providing particularly deep coverage over genes that are relevant to cancer development, inherited disorders, or other such conditions. This deeper coverage provides a higher level of confidence that detected variants are true hits and not artifacts of the sequencing process. The usefulness of a panel thus depends on where this deeper coverage is focused.
Twist Exome 2.0 provides deep coverage over all genes that have been linked to clinical phenotypes, enabling translational research on a wide range of conditions. It can be used to study known or potential variants associated with germline cancers, rare diseases, and inherited diseases, providing researchers with a windfall of high-quality, informative data. And, the data generated by the Twist Exome 2.0 panel extends beyond the exome. Spiked coverage over noncoding regions that are known to carry pathogenic or likely pathogenic variants enhances the Twist Exome 2.0 panel’s discovery power.
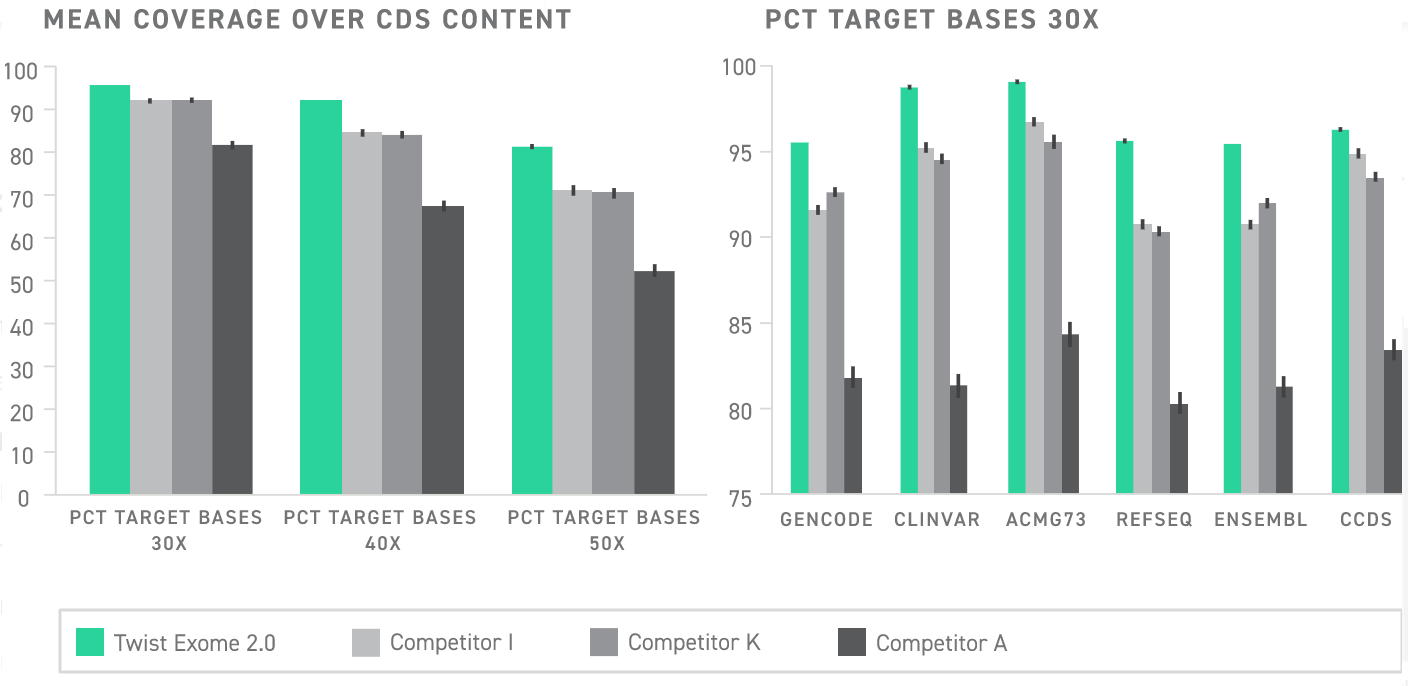
Coverage of databases relevant to clinical research
Determining where to provide deep coverage for whole exome sequencing can be challenging. Guidance can be found in databases like ClinVar and Genecode which provide a wealth of genomic annotations identifying variants and loci within the genome that are linked to clinical phenotypes. These high-impact regions are invaluable for translational research because they can shine light on the mechanisms driving disease pathology, reveal potential biomarkers, and may influence the effectiveness of prospective therapeutics. Because of this, researchers benefit from exome panels that maximize coverage over these databases.
Twist Exome 2.0 offers complete coverage over multiple of these resources, including ClinVar and the ACMG73. Relative to all other exomes currently on the market, Twist Exome 2.0 provides the best coverage over ClinVar, ACMG73, and Refseq.
By providing this broad and deep coverage, Twist Exome 2.0 opens the door for researchers to build a more comprehensive picture of both rare and common genetic variation, and how these influence a wide range of clinical phenotypes.
In short, there’s no need to compromise on coverage over performance. Twist Exome 2.0 provides best-in-class coverage with best-in-class performance.
Efficiency through uniformity and on-target rate
In addition to broad coverage, Twist Exome 2.0 offers unparalleled efficiency through uniformity and a high on-target rate.
Uniformity refers to how even target capture occurs. Without uniformity, target enrichment is more likely to be biased toward the capture of one sequence over another. In effect, this means many more rounds of sequencing would be needed to ensure that poorly captured targets are adequately represented. This is summarized as a fold-80 score, which describes how much over-sequencing must be performed to bring 80% of the targeted sequences up to mean coverage. A lower fold-80 score indicates a more uniform representation across targets.
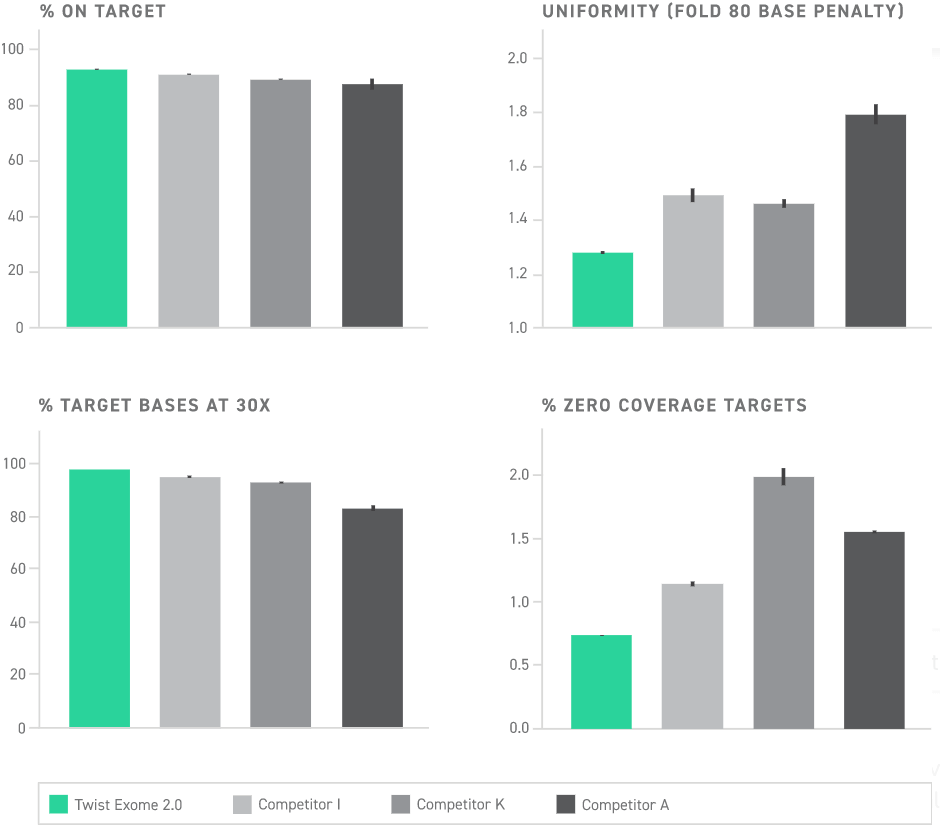
When the fold-80 score is high, you begin to accumulate costly, wasted sequencing reads for the well-captured targets. Therefore, lowering the fold-80 score can be cost-effective as you reduce the amount of sequencing needed to assess each target adequately.
Similarly, a low on-target rate can result in wasted sequencing. On-target rates describe how often the desired genetic sequence is captured by the panel (the inverse of this is off-target rates, how often undesired sequences are captured).
With the lowest fold-80 score and highest on-target rate among commercially available exomes, Twist Exome 2.0 is industry-leading in its efficiency—achieving 50x coverage over 80% of its target in a protocol that is twice as fast as its top competitors. This translates to having twice as much sequencing currency which can be spent going deeper for target genes, or sequencing twice as many samples per sequencing run.
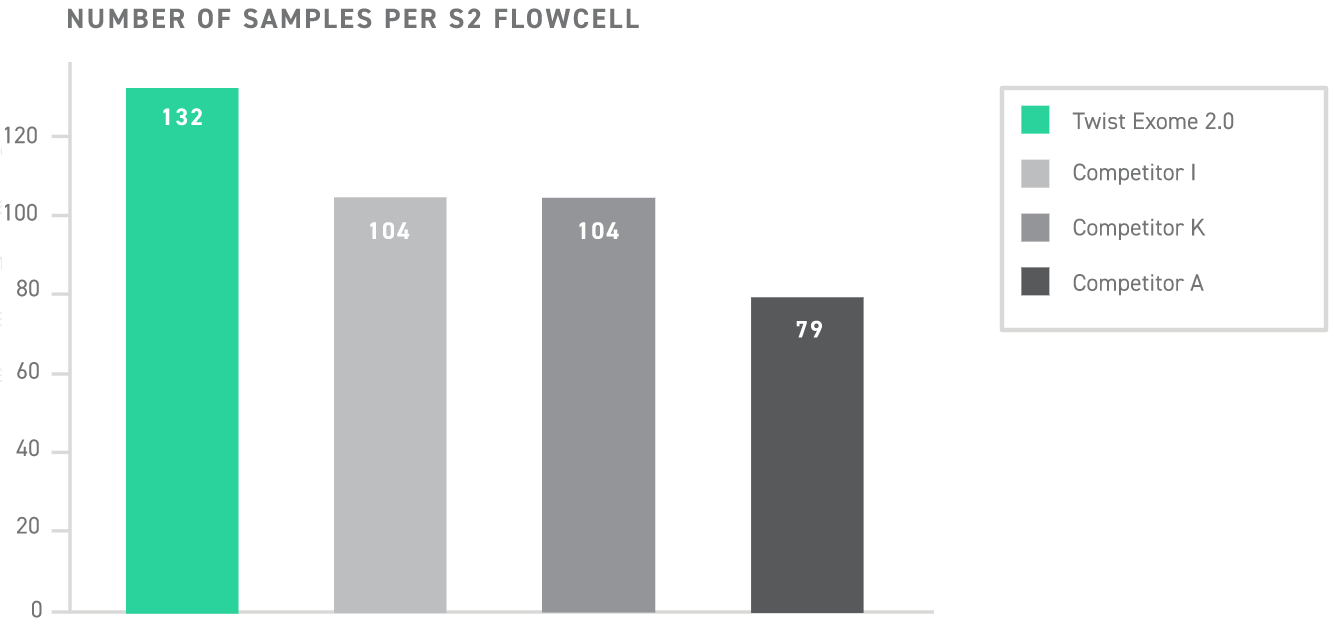
A new standard in WES
The Twist Exome 2.0 panel is best-in-class because it brings together broad coverage with unparalleled efficiency, enabling researchers to go deeper and sequence more samples per run. In short, this panel is designed to give you the type of high-quality data it takes to find answers and detect the unexpected.
The Twist Exome 2.0 panel is for Research Use Only. Not for use in Diagnostic Procedures.
What did you think?
Like
Dislike
Love
Surprised
Interesting