Just a stone’s throw away from the Tennessee River in Huntsville, Alabama is the HudsonAlpha Institute for Biotechnology. HudsonAlpha is a non-profit institution whose vision is “To leverage the synergy between discovery, education, medicine, and economic development in genomic sciences to improve the human condition around the globe.”
HudsonAlpha is a world leader in DNA sequencing and the hub for all sequencing activity within the institute is the Genomic Services Laboratory (GSL), run by Shawn Levy, PhD. The GSL houses a state-of-the art suite of Illumina sequencers, a dedicated bioinformatics group, and provides sequencing services at a global scale, completing about 4,500 projects. Over the last year, Dr. Levy and HudsonAlpha worked with Twist Bioscience to conduct a successful pilot project that shows the benefits of Twist Bioscience’s Target Enrichment Solutions for the next-generation sequencing (NGS) of Formalin-Fixed, Paraffin-Embedded (FFPE) tissue samples. This project’s findings were presented at AGBT 2018 in Orlando, FL. This work promises reduced costs and higher flexibility of NGS pipelines in clinical diagnostics and clinical research, and we will dive deeper into it here.
FFPE samples are small, preserved patient biopsies used for either patient diagnostics or future research into health and disease. Composed of a solution of formaldehyde and methanol in water, formalin is known mostly for its use in embalming. It creates a strong chemical bond between the amino acids of protein structures, making it an exceptional preserving agent for biological tissue. For long term storage, formalin-preserved samples are embedded in immunohistochemistry-grade paraffin. FFPE preserved tissues can be stored at room temperature for decades.
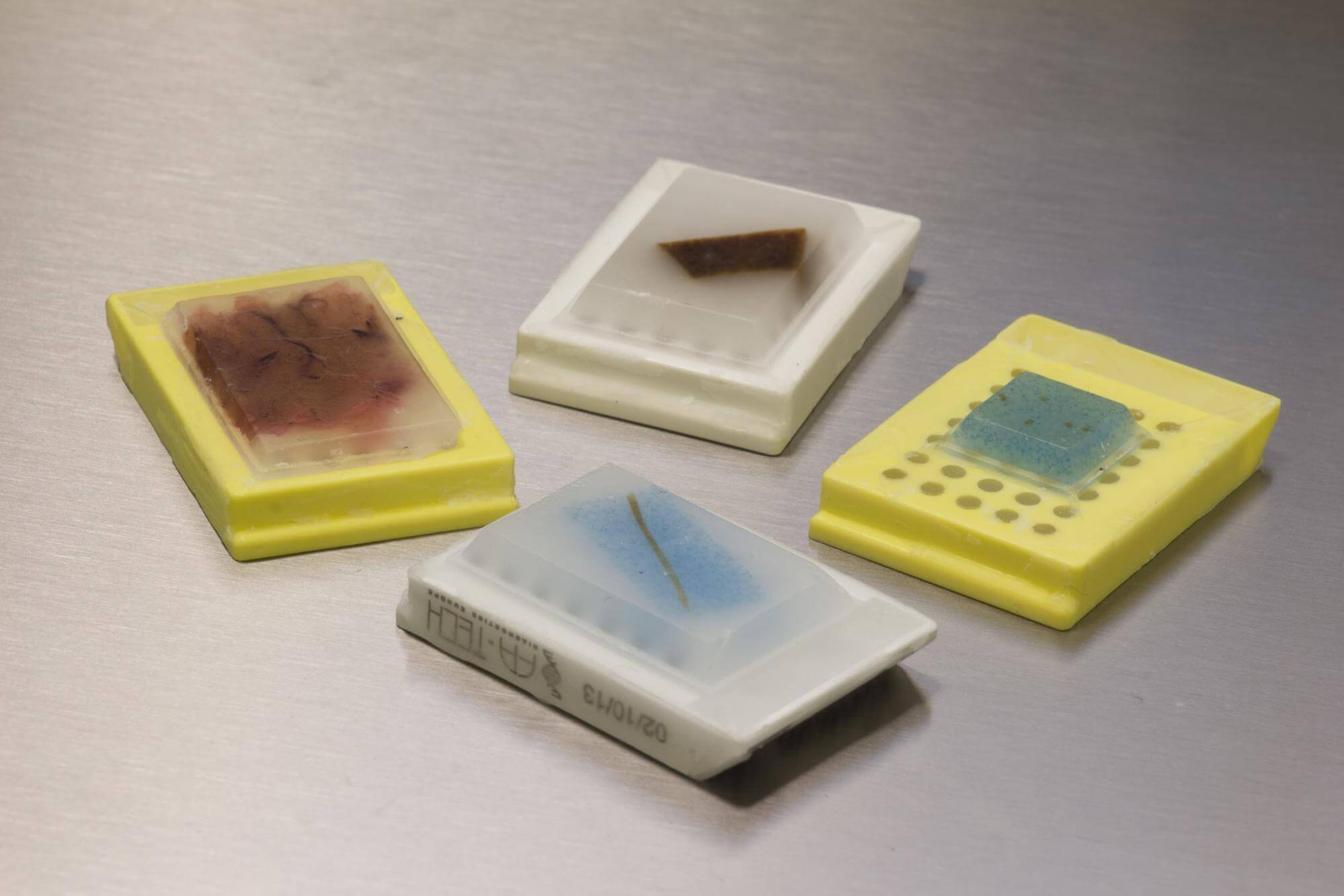

FFPE tissue samples are stored on cassettes.
FFPE samples have long been the standard storage and preparation method for tissues used in immunohistochemistry (the study of protein presence and position in a cell or tissue using fluorescent antibodies), and histopathology (the microscopic study of diseased tissue). Millions of tissue samples are archived in hospitals, tissue banks and research centers. FFPE preservation is pivotal to ongoing research on many diseases, including cancer and Alzheimer’s.
Importantly, where there is tissue, there is DNA. And where there is DNA from diseased tissue, there is an opportunity to learn about the genetics of disease through genomics and next-generation sequencing. There is a treasure trove of genetic information waiting to be sequenced from the diverse archived FFPE samples, which could provide vital information in the study of disease. Over the last 15 years, geneticists have been increasing their focus on the sequencing of DNA held within FFPE preserved tissues— an area where HudsonAlpha excels.
Unfortunately, extracting genetic information from FFPE samples isn’t simple. To learn more about the process and the study, we spoke to Dr. Levy in interview. He explained the challenges faced in the sequencing process: “A big problem is the quality of nucleic acid sample that comes out; it is really damaged during fixation.” When a cell is fixed in formalin, its genome is blown apart. Formalin and organic molecules react readily, so when it comes into contact with DNA, it causes double stranded breaks and cross linking. “Base modifications or other DNA damage events we see in the sample make the sequencing more difficult,” added Dr. Levy.
As a result, FFPE samples return very poor quality sequencing data that is often useless or unreliable. However, if enough DNA is sequenced from a sample, useful genetic information can still be compiled. With the advent of low cost next-generation sequencing, the study of FFPE samples is becoming increasingly viable. Additionally, certain preparation and storage protocols are more likely to produce higher quality data.
“While sample age is a factor, the fixation protocol and storage method are the main problem,” explained Dr. Levy. “For example, if we have access to large enough samples, we can get 30-40 microns into the tissue and it has not been exposed to air. We find old samples that are large enough that they can produce higher quality data than smaller samples that have just been fixed - in those samples we often see a lot of damage.” This kind of damage reduces the amount of usable material for downstream processing and analyses. Additionally, each FFPE sample is finite, often without options to retrieve more tissue from the same patient if experiments fail. If a FFPE sample is precious, then it's essential that high-quality data can be retrieved.
Difficulties in sequencing FFPE tissue samples require specialized sample preparation and sequencing protocols in order to obtain meaningful data. One powerful approach is targeted sequencing. Targeted sequencing leverages DNA-based hybridization probes that capture and isolate only the sequences of interest within the sample. For example, around 85% of rare genetic diseases manifest in humans from mutations in our exome.
The exome is the part of the genome that encodes proteins, accounting for less than 2% of our entire genetic material. Targeting only sequences of interest within a sample allows researchers to direct resources toward obtaining the read depth and coverage required to make the correct conclusions from lower quality FFPE data. This is especially salient for studies of cancer samples, where disease causing mutations may be “rare” in the context of the entire sample. In such complex cases, a high-read depth (>500x) is required. This ensures that when the sequencing data is aligned to a reference, disease causing mutations can be separated from background mutations caused by the FFPE preparation process.
To advance HudsonAlpha’s research on FFPE samples, Dr. Levy was one of the first customers to test Twist Bioscience’s Human Core Exome Kit. Twist Bioscience is a leader in high-throughput DNA synthesis, allowing Twist to offer highly uniform target enrichment solutions. To assess performance, Dr. Levy's team used a diverse set of FFPE samples with varying quality which they captured with the Twist exome and subsequently sequenced at 500x.

Shawn Levy, PhD runs the Genomic Services Laboratory at HudsonAlpha, and was one of the first customers to test Twist Bioscience’s NGS Target Enrichment Solutions
In previous experiments with other target capture solutions, HudsonAlpha found that when deep sequencing FFPE preserved tissue, around 92% of targets could be sequenced to 30x. However, when running the same experiment with Twist Bioscience’s Human Core Exome Kit, >99.5% of all targeted samples were covered with 30 or more reads providing strong support for variants detected. Not only that, but Twist’s Human Core Exome Kit was found to have a lower duplication rate compared to other kits on the market.
Dr. Levy was very positive about his experience with Twist’s Human Core Exome Kit: “The preparation of the library itself was very easy, with many of the key points in the protocol were ‘addition only’ steps,” he said. “Downstream, this makes the protocol easy for us to standardize, and in turn simple to automate.” He added that, “The sample input requirement per experiment is significantly lower than other kits that we have worked with, this is really important when were dealing with the tiny, finite amount of FFPE preserved sample.”
For HudsonAlpha, these benefits have immediate positive potential for their sequencing workflows. “Data acquisition is currently a bottleneck, and it needs to be streamlined,” Dr. Levy said. “Anything that saves time is incredibly valuable to clinical diagnostics, as time is a precious and expensive resource, especially for clinical diagnostics of cancer and infectious disease.”
For the future of molecular diagnostics, Dr. Levy expressed that he felt “Targeted sequencing will lead the way for reimbursable diagnostics in the short-term, for both whole genome sequencing and whole exome sequencing.” He explained: “Exome sequencing experiments create huge volumes of data, and we aren’t putting that data to use as efficiently as we could. Broad scale phenotyping will be the next frontier, and exome sequencing gives us the volume of data we need, now we just need to leverage it.”
From our conversation with Dr. Levy, it seems that we are on the cusp of an inflection point in molecular diagnostics, now that target capture is highly efficient for even the most challenging of samples. It's possible to produce the high volume data required using tools produced by next-generation synthesis providers like Twist Bioscience, even when “difficult” samples like FFPE are to be sequenced. This is a significant development for NGS. By honing in on exactly the genetic material of interest with pinpoint accuracy using tools like Twist Bioscience’s NGS Target Enrichment Solutions, cost-per-experiment can be offset with the volume of experiments that yield valuable results. This effect is maintained beyond clinical diagnostics to clinical research, meaning contemporary advancements in DNA synthesis are pivotal to the future of next-generation sequencing.
Subscribe to our blog
- Terms & Conditions
- Policy on Unsolicited Submissions of Information
- Privacy Policy
- Regulatory and Quality Information
- Security
- Payment Information
-
© 2025 Twist Bioscience. All rights reserved.