Twist Bioscience
December 12, 2017
min read
Written by Emily Leproust
Originally published on TechCrunch Sep 18, 2015
Originally published on TechCrunch Sep 18, 2015
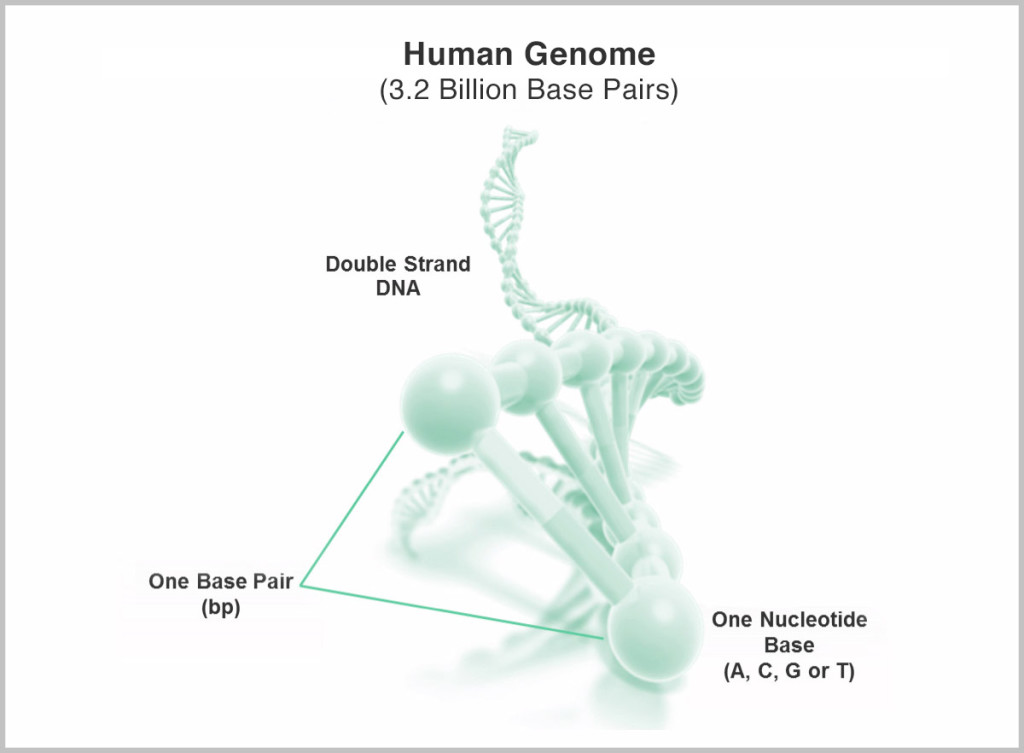
The “$1,000 genome” has been one of the most visible tech goals in the last 10 years, even sporting its own XPrize for a short time. It refers to the ability to “sequence,” or de-code, the total genetic information encoded in the DNA (genome) of a single human, at a cost of $1,000. It’s a daunting task, given that the genome is made up of 3 billion base pairs whose exact sequence needs to be determined.
But why $1,000? Many experts feel this is the cost at which it becomes feasible to sequence human genomes on a large scale. The data could then be used to improve clinical diagnosis and treatment of rare and new disorders suspected of having genetic causes. Illumina hit the $1,000 mark in January of 2014.
Their instruments are capable of sequencing 18,000 human genomes per year. Many individual genomes have now been sequenced and have proved invaluable for finding causal genetic mutations for such diseases as schizophrenia and autism. The sequencing of the genome has played a role in personalized genomics, genetic testing and forensic science, just to name a few. This very significant breakthrough in DNA sequencing capability heralds a new era in medicine.
Sequencing DNA can be thought of as the “reading” step, which gives us the information that codes for life. The next step is to “write” new genetic information by synthesizing genes. As researchers, whether the field is agricultural biology, medicine, academia or industrial chemical, the cycle of research is the same — design, build, test — and it repeats until a solution is identified.
This process continues as it has for many years, but there is a bottleneck in the synthesis of genes. It is a time-consuming and expensive proposition, and researchers in a variety of industries are frustrated with the existing process. That’s where synthetic biology (synbio for short) comes in.
Over the last three decades, cloning has been the method of writing genetic material de novo (DNA, RNA and proteins) in all industries using biological materials. This includes industrial chemistry, agricultural biotechnology, medicine and academia. Rapid and affordable access to large quantities of genetic material will drive advances in synthetic biology, which promises to improve health, help clean up the environment, feed the world, and create new industries and jobs.
Current DNA synthesis technologies cannot meet research demand. As an industry, we need to further reduce the financial costs and turnaround times. Today, obtaining 100 to 1,000 genes can take weeks or even months from conventional suppliers, with a cost of $400 per 2000 bp gene ($0.20 per base pair) or more. These costs are prohibitively high, particularly for academic users. They can make their own DNA at a significantly reduced monetary cost, but it can take weeks of tedious labor, which is a cost of another kind.
Several companies provide synthesis of oligonucleotides (small pieces of DNA) and genes, but most of them use the same approach that has been around for many years. These traditional synthesis methods use one 96-well plastic plate to make one gene. A different oligonucleotide (oligo) is synthesized in each well using chemistry developed 30 years ago.
The 96 oligos are then removed from the plate and assembled into one gene using enzymes to amplify each one and then connect them into a long string, a process that is both slow and tedious. The gene is then inserted into bacteria (cloned) to produce billions of copies of the gene and make them available for further use. This approach was revolutionary in 1985 but is not sufficient to provide the volume and speed needed to meet the needs of modern synthetic biology.
Companies using conventional gene synthesis approaches include Life Technologies (GeneArt), Integrated DNA Technologies (SGI-DNA), DNA 2.0 andGenScript. Costs can range from $.25 per base pair (bp) to as much as $.60 per bp ($1,200 for a 2000 bp gene), and the multi-step process can lead to long delivery times of a few weeks or even longer for large genes.
Gen9, a Cambridge MA startup, takes a somewhat different approach, using highly parallel synthesis on a semiconductor platform as an attempt to increase throughput. The oligonucleotides used to assemble the genes are made in “pools” that are then released from the chip, de-pooled, amplified and assembled into genes using enzymes. My company, Twist Bioscience, utilizes a platform for high-density synthesis of genes on silicon wafers. Using this technology, DNA oligo synthesis and gene assembly steps occur right on silicon.
Getting back to the $1,000 genome, the equivalent of this cost in the world of DNA writing is $0.02 per base pair. That’s the point at which researchers can significantly scale experiments, and will no longer be limited by the cost of DNA. With the advances in DNA synthesis, this goal is within reach—in much less time than the decade it took to achieve the $1,000 genome.
The medical and life sciences stand on the threshold of an era that will usher in a more in-depth understanding of biological and disease processes at a molecular level, made possible by synthetic biology. The practical applications of this new capability will have profound positive effects on healthcare, the environment, and the economy.
Get the latest by subscribing to our blog
Get the latest
Subscribe to our blog
INTEGRATIONS
Technology
Resources
Partnerships
- Terms & Conditions
- Policy on Unsolicited Submissions of Information
- Privacy Policy
- Regulatory and Quality Information
- Security
- Payment Information
-
© 2025 Twist Bioscience. All rights reserved.